Semiparametric Causal Inference in StochTree
Source:vignettes/ReparameterizedCausalInference.Rmd
ReparameterizedCausalInference.RmdThe classic BCF model of Hahn, Murray, and Carvalho (2020) is defined as where defines a BART model with trees and split prior parameters and .
The authors noted that separating estimation / regularization of a control function, , and a CATE function, , can give better estimation error and interval coverage in settings with strong confounding and treatment effect moderation.
stochtree now defaults to a slight modification of this
model, with the treatment effect function decomposed into parametric and
nonparametric components
where
takes the place of the
forest term in the original BCF model. This decomposition allows the
forest term to focus on capturing heterogeneity “offsets” to a
parametric model of homogeneous treatment effects.
Below we demonstrate the advantages of this “reparameterization” of BCF on a synthetic dataset.
First, we load the necessary libraries
We set a seed for reproducibility
random_seed <- 1234
set.seed(random_seed)Binary Treatment with Homogeneous Treatment Effect
Consider the following data generating process
Simulation
We draw from the DGP defined above
n <- 500
p <- 20
snr <- 2
X <- matrix(runif(n * p), n, p)
mu_x <- 2 * sin(2 * pi * X[, 1]) - 2 * (2 * X[, 3] - 1)
tau_x <- 5
pi_x <- pnorm(mu_x / 4)
Z <- rbinom(n, 1, pi_x)
E_XZ <- mu_x + Z * tau_x
sigma_true <- sd(E_XZ) / snr
y <- E_XZ + rnorm(n, 0, 1) * sigma_trueAnd split data into test and train sets
test_set_pct <- 0.5
n_test <- round(test_set_pct * n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- X[test_inds, ]
X_train <- X[train_inds, ]
pi_test <- pi_x[test_inds]
pi_train <- pi_x[train_inds]
Z_test <- Z[test_inds]
Z_train <- Z[train_inds]
y_test <- y[test_inds]
y_train <- y[train_inds]
mu_test <- mu_x[test_inds]
mu_train <- mu_x[train_inds]
tau_test <- tau_x[test_inds]
tau_train <- tau_x[train_inds]Sampling and Analysis
Classic BCF Model
We first simulate from the classic BCF model with no parametric treatment effect term
num_gfr <- 0
num_burnin <- 1000
num_mcmc <- 500
general_params <- list(
adaptive_coding = TRUE,
num_chains = 4,
random_seed = random_seed,
num_threads = 1
)
num_trees_tau <- 50
treatment_effect_forest_params <- list(
num_trees = num_trees_tau,
sample_intercept = FALSE,
sigma2_leaf_init = 1 / num_trees_tau
)
bcf_model_classic <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)And we compare the posterior distribution of the ATE to its true value
cate_posterior_classic <- predict(
bcf_model_classic,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_classic <- colMeans(cate_posterior_classic)
hist(
ate_posterior_classic,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)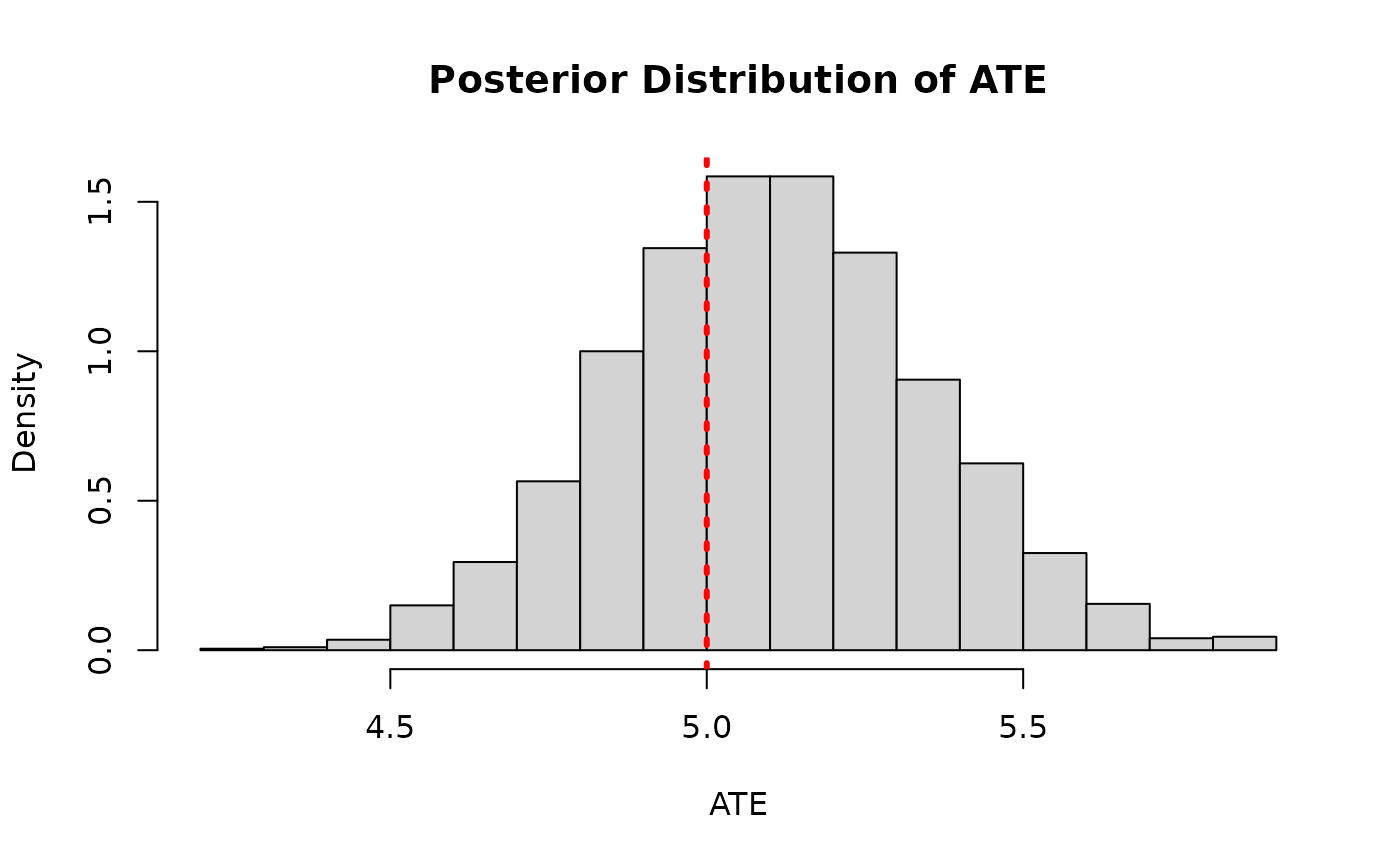
As a rough convergence check, we inspect the traceplot of the global error scale parameter,
sigma2_samples <- extractParameter(bcf_model_classic, "sigma2")
plot(
sigma2_samples,
type = "l",
main = "Traceplot of Sigma^2",
ylab = "Sigma^2",
xlab = "Iteration"
)
abline(h = sigma_true^2, col = "red", lty = 3, lwd = 3)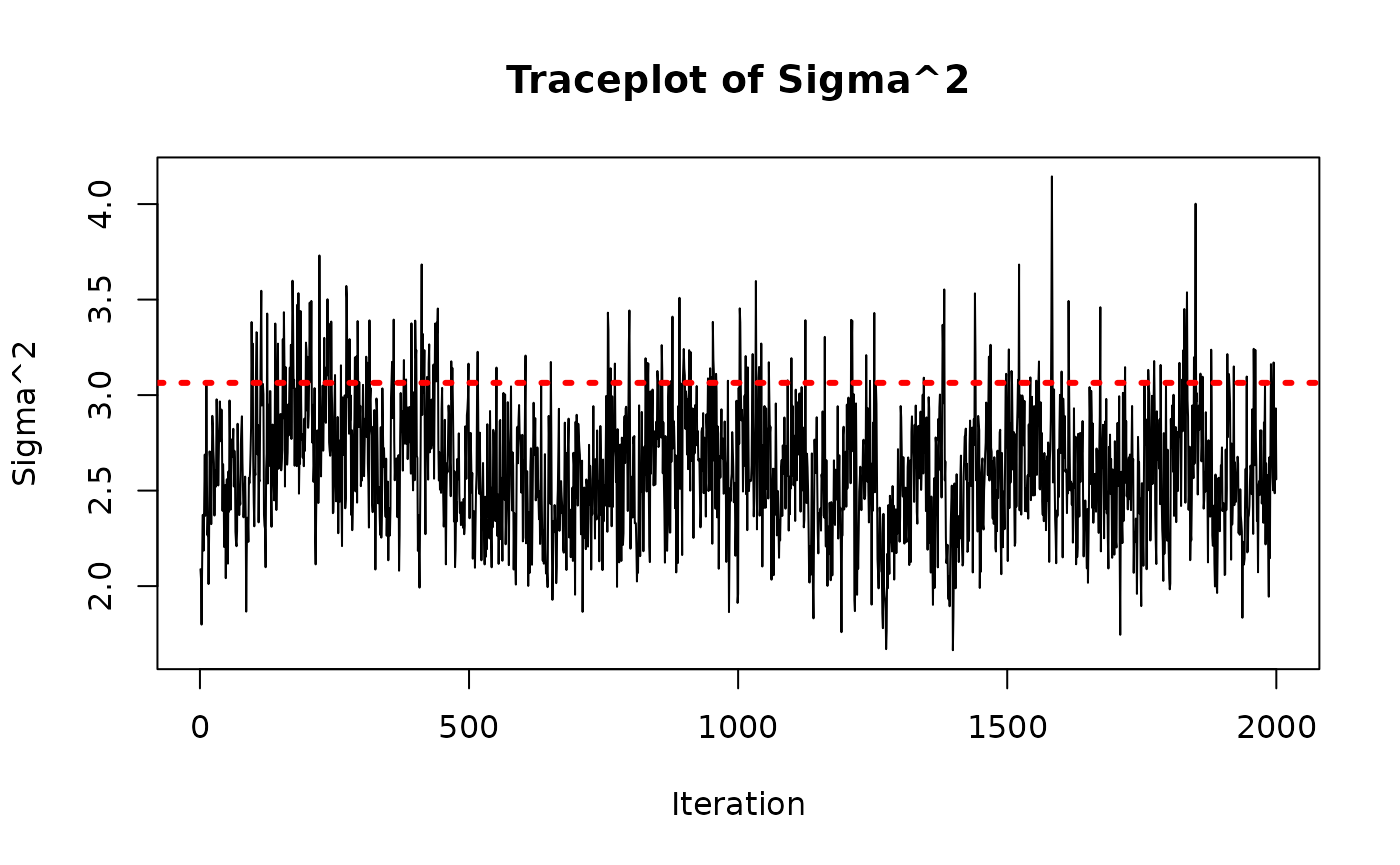
Reparameterized BCF Model
Now we fit the reparameterized model, regularizing the forest more heavily to account for the standard normal prior on the term.
num_trees_tau <- 50
general_params <- list(
adaptive_coding = FALSE,
num_chains = 4,
random_seed = random_seed,
num_threads = 1
)
treatment_effect_forest_params <- list(
num_trees = num_trees_tau,
sample_intercept = TRUE,
sigma2_leaf_init = 0.25 / num_trees_tau,
tau_0_prior_var = 1
)
bcf_model_reparam <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)And we compare the posterior distribution of the ATE to its true value
cate_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_reparam <- colMeans(cate_posterior_reparam)
hist(
ate_posterior_reparam,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)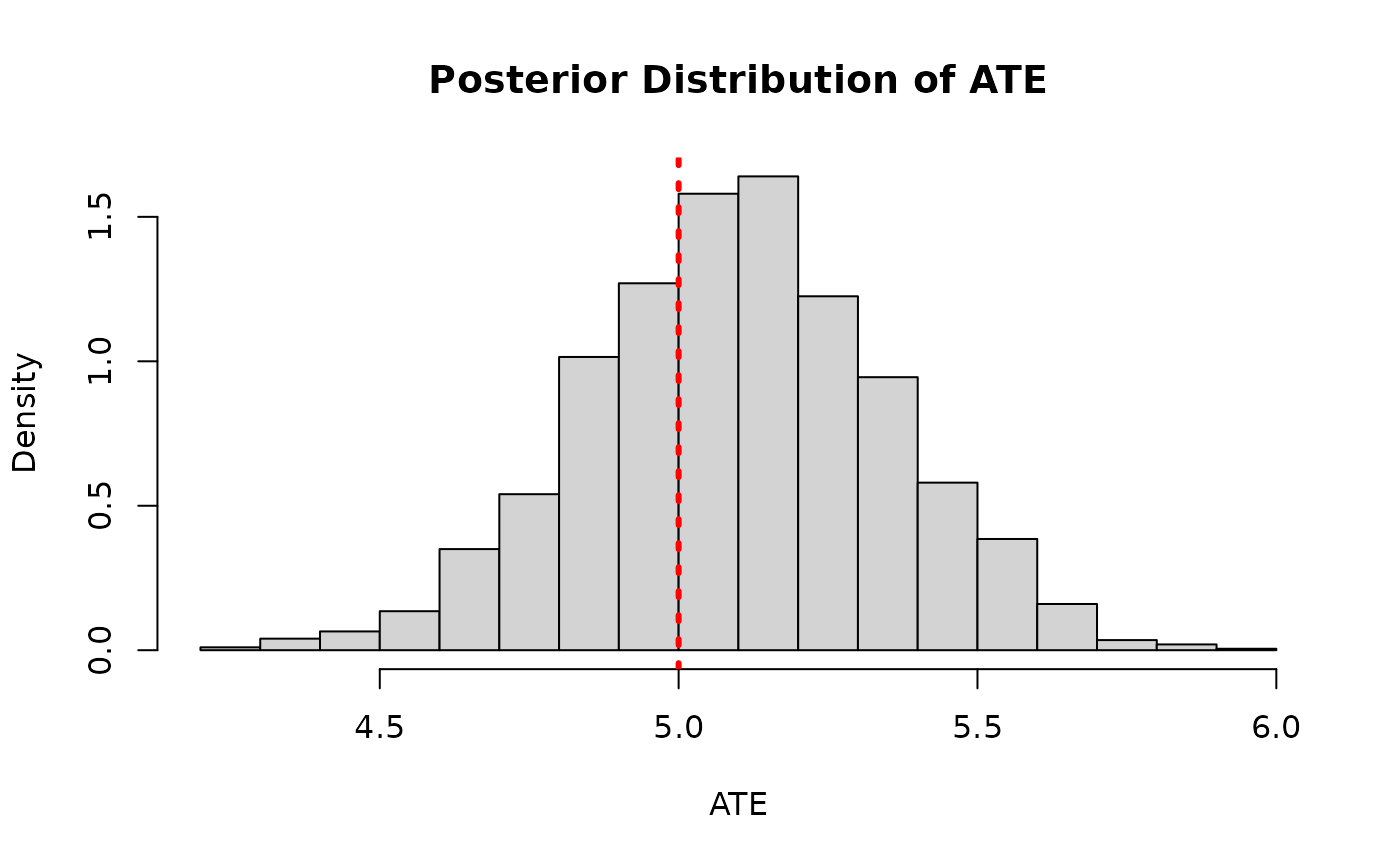
As above, we check convergence by inspecting the traceplot of the global error scale parameter,
sigma2_samples <- extractParameter(bcf_model_reparam, "sigma2")
plot(
sigma2_samples,
type = "l",
main = "Traceplot of Sigma^2",
ylab = "Sigma^2",
xlab = "Iteration"
)
abline(h = sigma_true^2, col = "red", lty = 3, lwd = 3)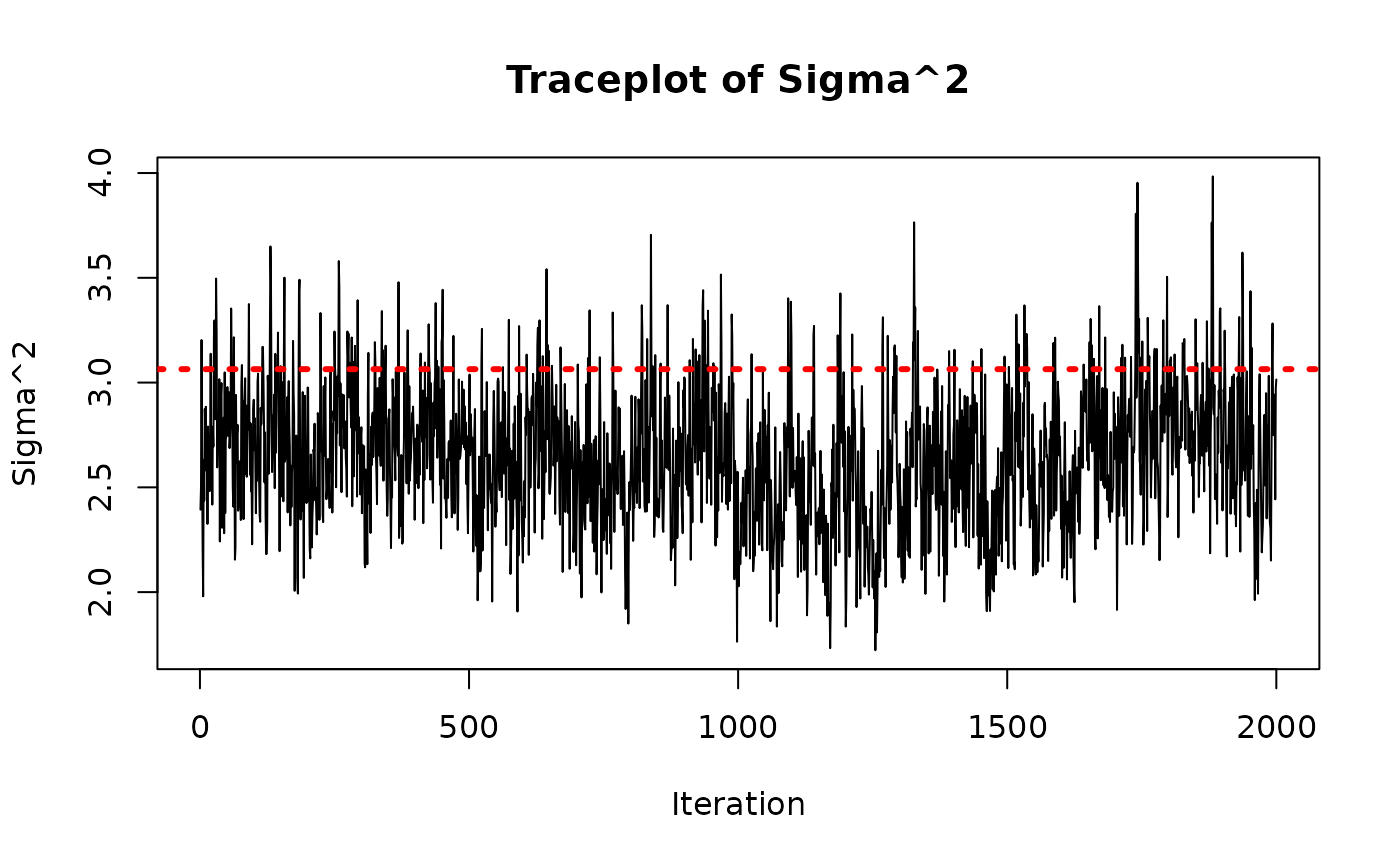
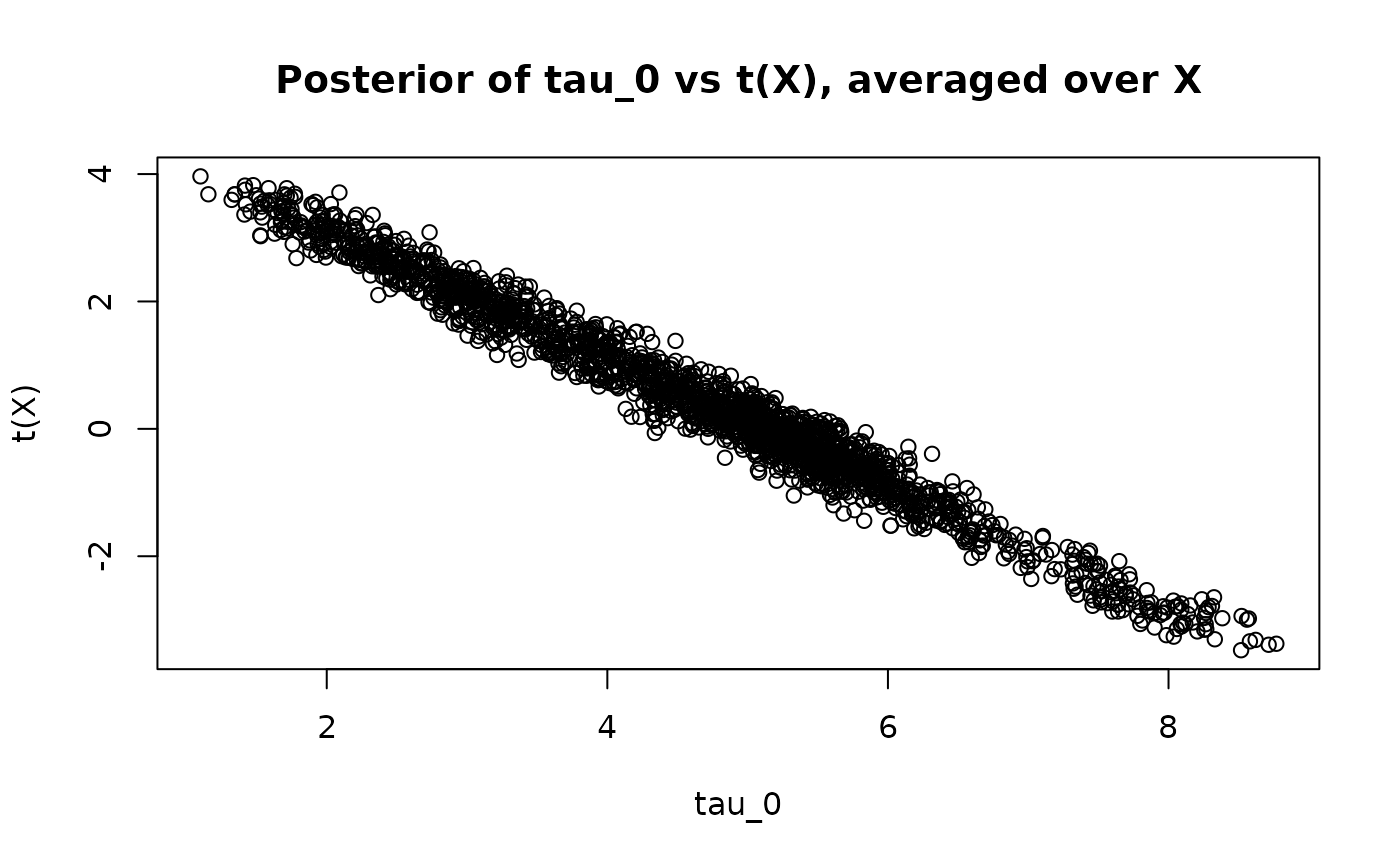
Since is not constrained to sum to 0, the parameter does not identify the ATE. We can see this by averaging each posterior draw of over the test set and comparing the posterior point estimates and .
tau_0_posterior <- extractParameter(bcf_model_reparam, "tau_0")
t_x_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "tau"
)
t_x_posterior_reparam <- colMeans(t_x_posterior_reparam)
plot(
tau_0_posterior,
t_x_posterior_reparam,
xlab = "tau_0",
ylab = "t(X)",
main = "Posterior of tau_0 vs t(X), averaged over X"
)
While stochtree does not currently support constraining
to sum to 0 over the training set, we can more heavily regularize
so that its values are much closer to zero. Using a single tree with a
very small leaf scale effectively collapses the forest to a constant
near zero, making
the primary vehicle for the treatment effect.
general_params <- list(
adaptive_coding = FALSE,
num_chains = 4,
random_seed = random_seed,
num_threads = 1
)
treatment_effect_forest_params <- list(
num_trees = 1,
sample_intercept = TRUE,
sigma2_leaf_init = 1e-6,
tau_0_prior_var = 1
)
bcf_model_reparam <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)Again we plot the posterior distribution of the ATE
cate_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_reparam <- colMeans(cate_posterior_reparam)
hist(
ate_posterior_reparam,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)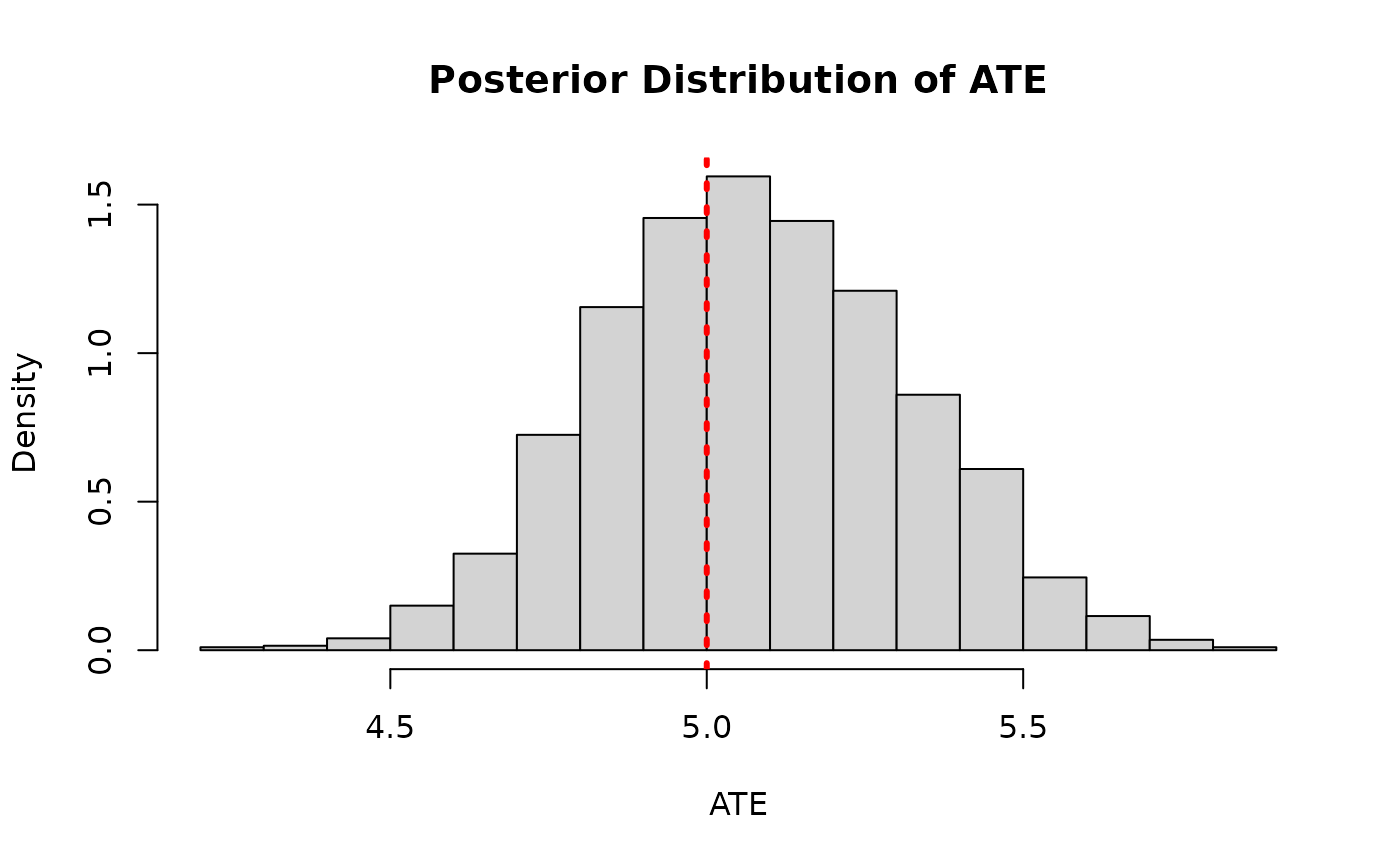
This time we see no correlation between the posterior and the (highly-regularized) posterior – more directly captures the majority of the ATE
tau_0_posterior <- extractParameter(bcf_model_reparam, "tau_0")
t_x_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "tau"
)
t_x_posterior_reparam <- colMeans(t_x_posterior_reparam)
plot(
tau_0_posterior,
t_x_posterior_reparam,
xlab = "tau_0",
ylab = "t(X)",
main = "Posterior of tau_0 vs t(X), averaged over X"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)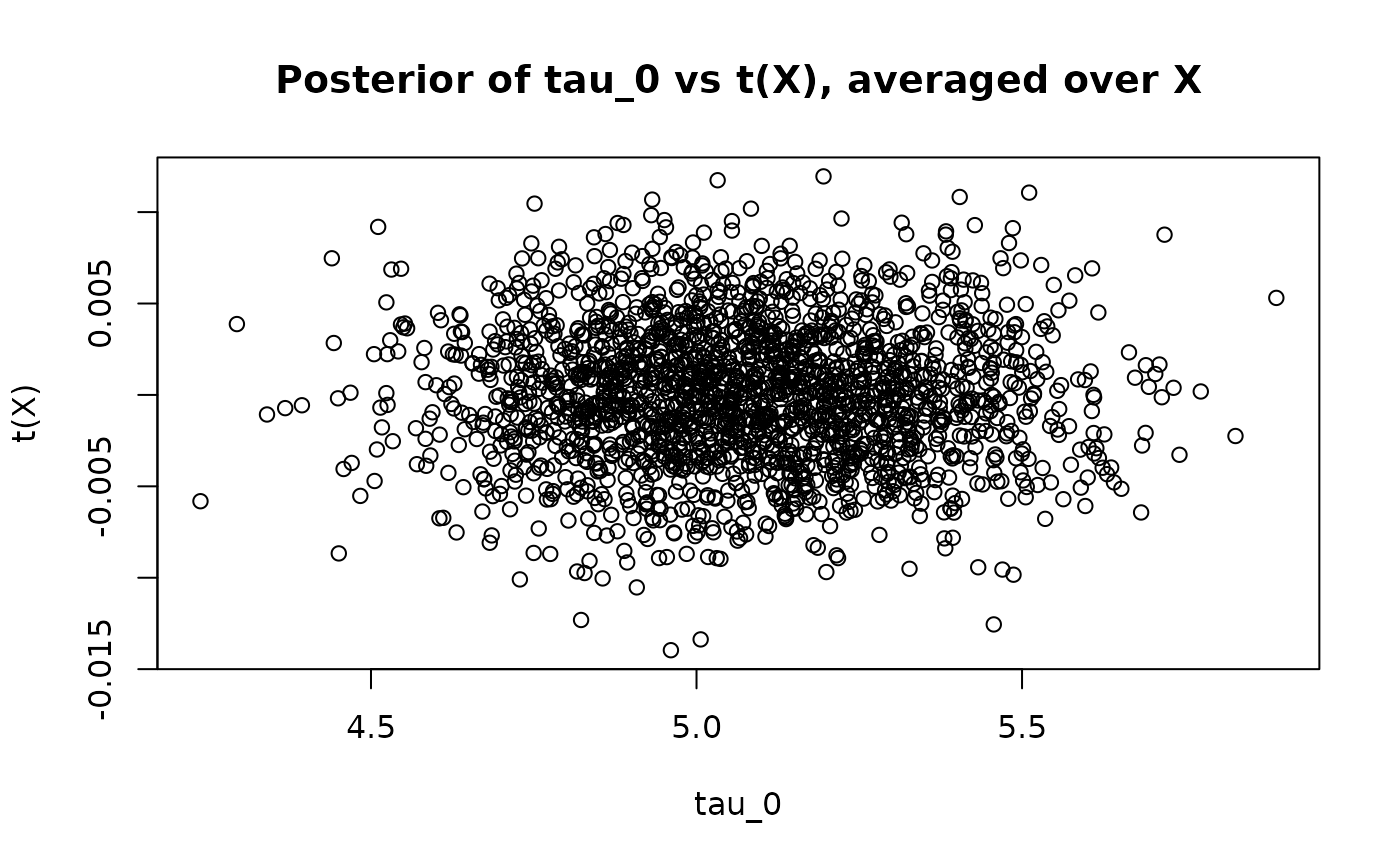
We can further regularize estimation of the ATE by reducing
general_params <- list(
adaptive_coding = FALSE,
num_chains = 4,
random_seed = random_seed,
num_threads = 1
)
treatment_effect_forest_params <- list(
num_trees = 1,
sample_intercept = TRUE,
sigma2_leaf_init = 1e-6,
tau_0_prior_var = 0.05
)
bcf_model_reparam <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)
cate_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_reparam <- colMeans(cate_posterior_reparam)
hist(
ate_posterior_reparam,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)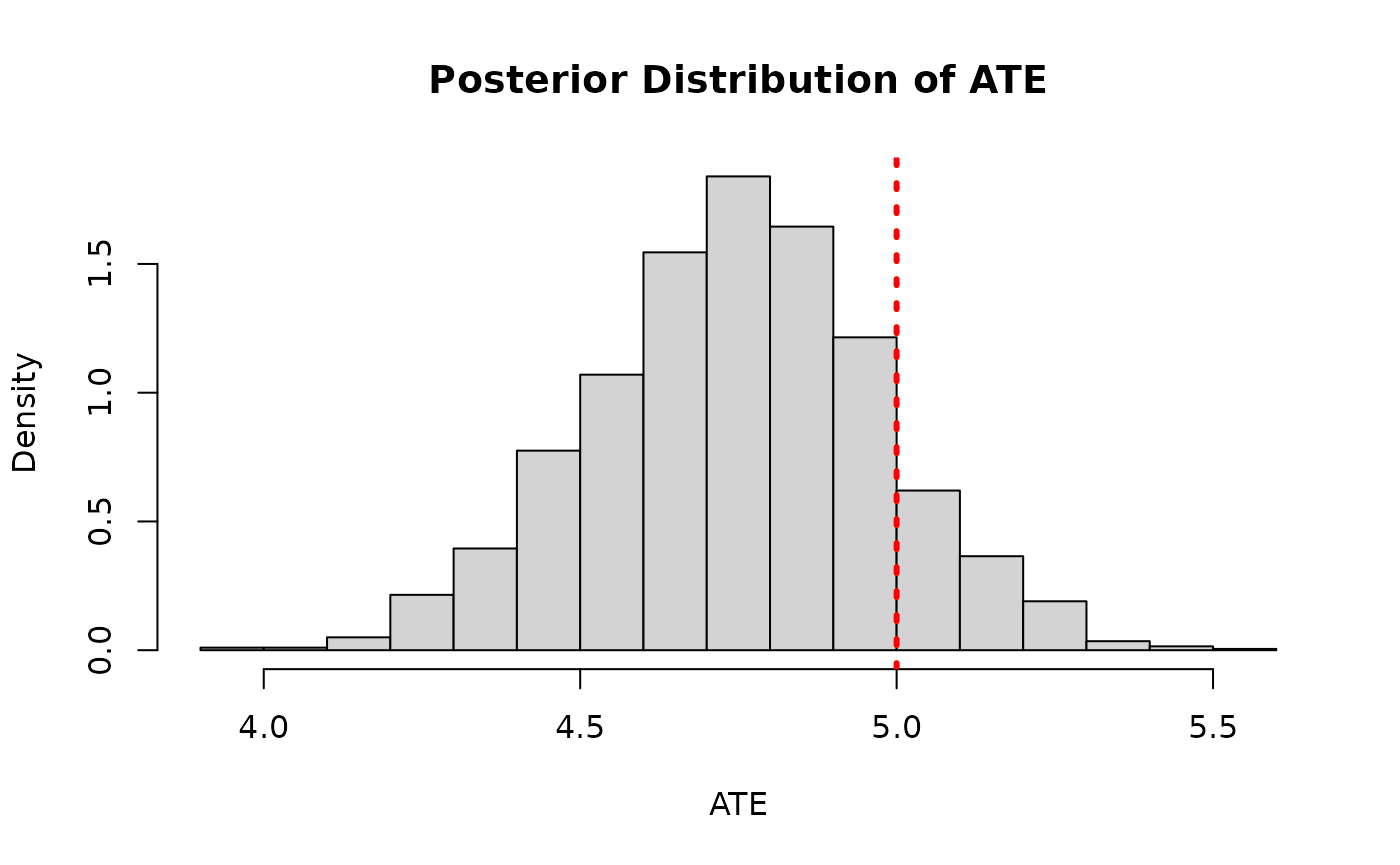
Continuous Treatment with Homogeneous Treatment Effect
The reparameterization generalizes naturally to continuous treatment. With a continuous , represents the marginal effect of a one-unit increase in , and captures the homogeneous component of that effect.
Consider the following data generating process:
Simulation
We draw from the DGP defined above
n <- 500
p <- 20
snr <- 2
X <- matrix(runif(n * p), n, p)
mu_x <- 2 * sin(2 * pi * X[, 1]) - 2 * (2 * X[, 3] - 1)
tau_x <- 2
pi_x <- mu_x / 8
Z <- pi_x + rnorm(n, 0, 1)
E_XZ <- mu_x + Z * tau_x
sigma_true <- sd(E_XZ) / snr
y <- E_XZ + rnorm(n, 0, 1) * sigma_trueAnd split data into test and train sets
test_inds <- sort(sample(1:n, round(0.5 * n), replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- X[test_inds, ]
X_train <- X[train_inds, ]
pi_test <- pi_x[test_inds]
pi_train <- pi_x[train_inds]
Z_test <- Z[test_inds]
Z_train <- Z[train_inds]
y_test <- y[test_inds]
y_train <- y[train_inds]Sampling and Analysis
Note that adaptive_coding must be FALSE for
continuous treatment, since the adaptive coding scheme is designed for
binary treatment.
Classic BCF Model
num_gfr <- 0
num_burnin <- 1000
num_mcmc <- 500
general_params <- list(
adaptive_coding = FALSE,
num_chains = 4,
random_seed = random_seed,
num_threads = 1
)
num_trees_tau <- 50
treatment_effect_forest_params <- list(
num_trees = num_trees_tau,
sample_intercept = FALSE,
sigma2_leaf_init = 1 / num_trees_tau
)
bcf_model_classic <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)We compare the posterior distribution of the ATE to its true value
cate_posterior_classic <- predict(
bcf_model_classic,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_classic <- colMeans(cate_posterior_classic)
hist(
ate_posterior_classic,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE (Classic BCF, Continuous Treatment)"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)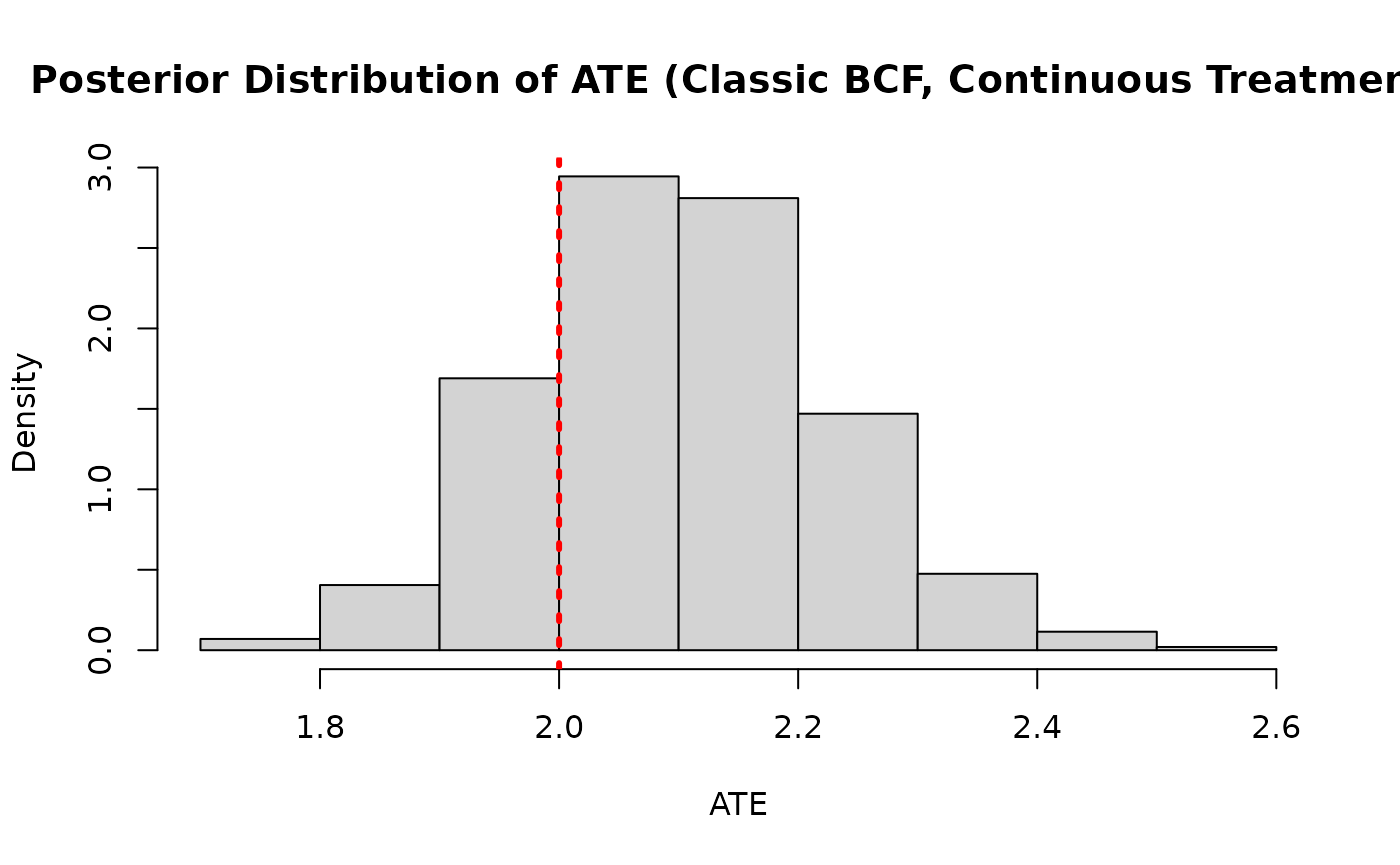
As a rough convergence check, we inspect the traceplot of
sigma2_samples <- extractParameter(bcf_model_classic, "sigma2")
plot(
sigma2_samples,
type = "l",
main = "Traceplot of Sigma^2 (Classic BCF, Continuous Treatment)",
ylab = "Sigma^2",
xlab = "Iteration"
)
abline(h = sigma_true^2, col = "red", lty = 3, lwd = 3)
Reparameterized BCF Model
treatment_effect_forest_params <- list(
num_trees = num_trees_tau,
sample_intercept = TRUE,
sigma2_leaf_init = 0.25 / num_trees_tau,
tau_0_prior_var = 1
)
bcf_model_reparam <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
treatment_effect_forest_params = treatment_effect_forest_params
)And we compare the posterior distribution of the ATE to its true value
cate_posterior_reparam <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "cate"
)
ate_posterior_reparam <- colMeans(cate_posterior_reparam)
hist(
ate_posterior_reparam,
freq = F,
xlab = "ATE",
ylab = "Density",
main = "Posterior Distribution of ATE (Reparameterized BCF, Continuous Treatment)"
)
abline(v = tau_x, col = "red", lty = 3, lwd = 3)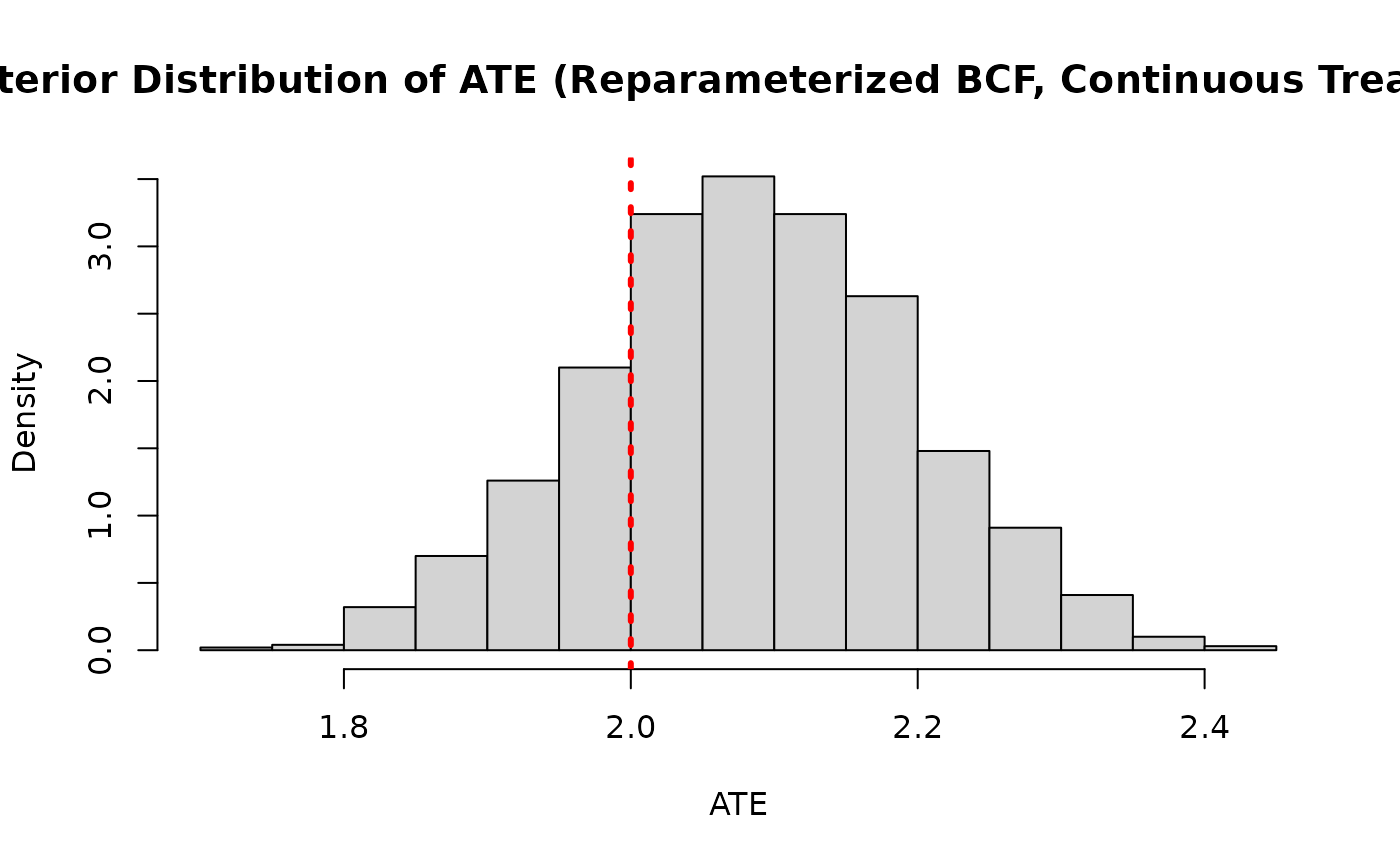
As above, we check convergence by inspecting the traceplot of
sigma2_samples <- extractParameter(bcf_model_reparam, "sigma2")
plot(
sigma2_samples,
type = "l",
main = "Traceplot of Sigma^2 (Reparameterized BCF, Continuous Treatment)",
ylab = "Sigma^2",
xlab = "Iteration"
)
abline(h = sigma_true^2, col = "red", lty = 3, lwd = 3)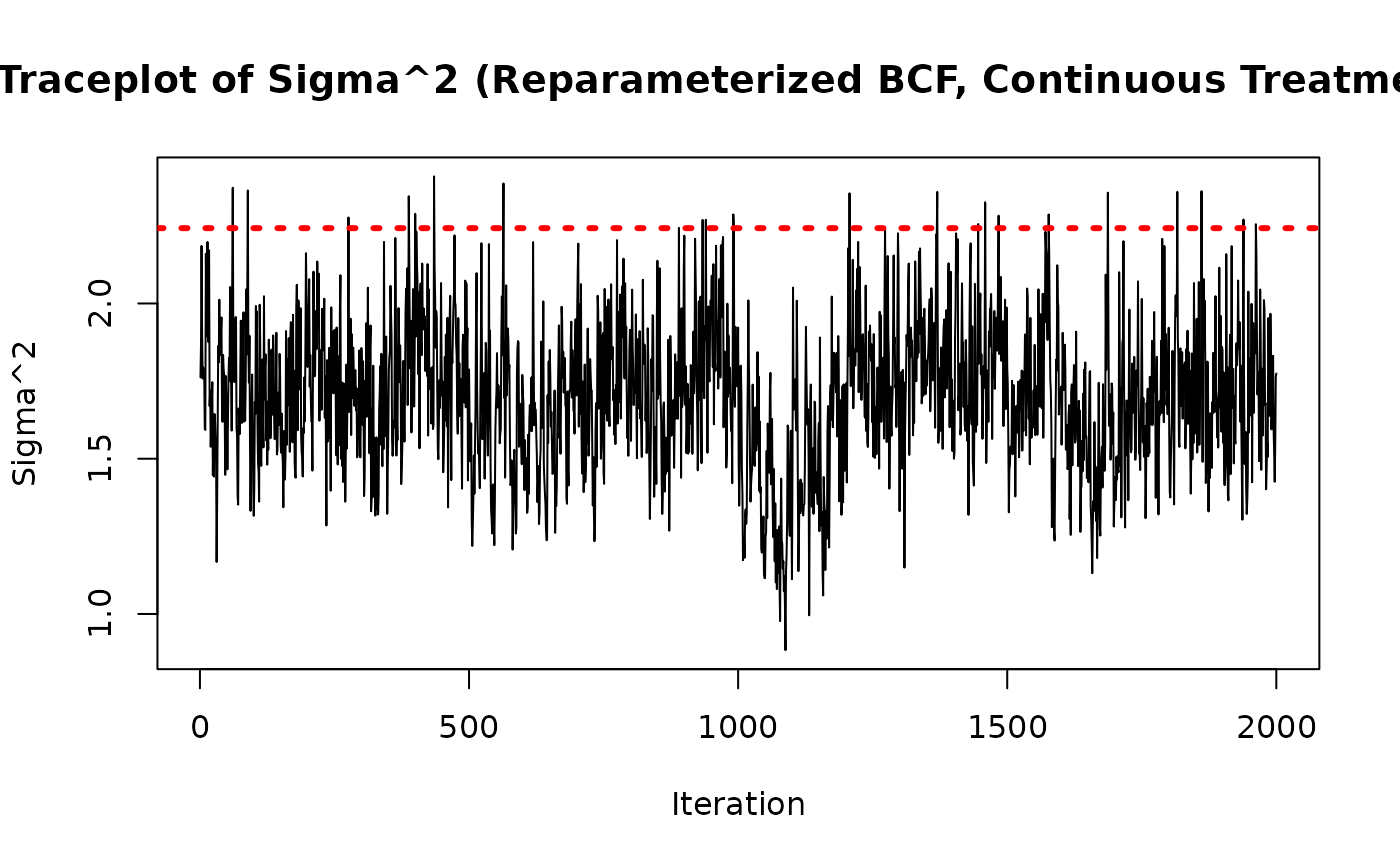
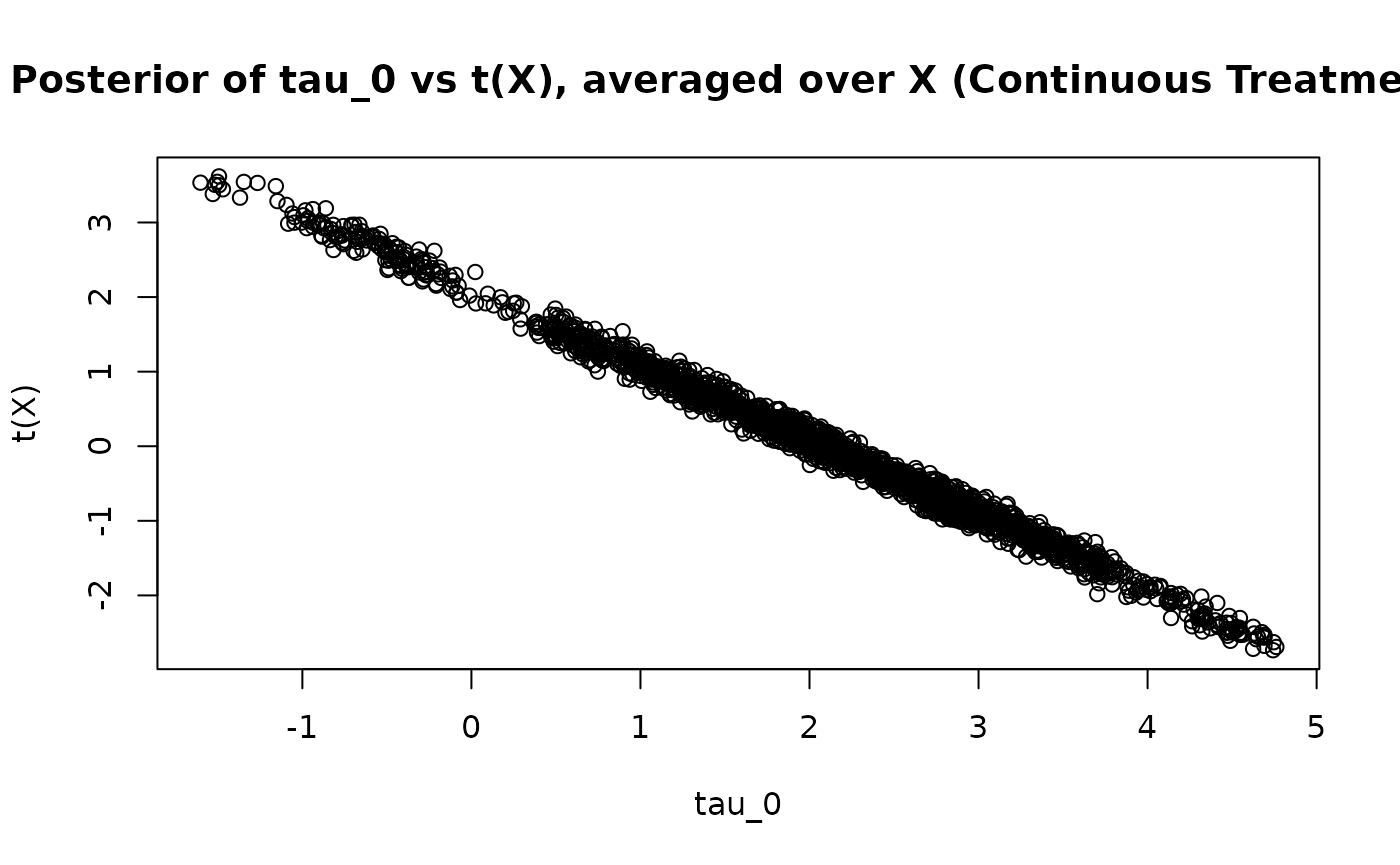
As in the binary treatment case, and are negatively correlated across posterior draws
tau_0_posterior <- extractParameter(bcf_model_reparam, "tau_0")
t_x_posterior <- predict(
bcf_model_reparam,
X = X_test,
Z = Z_test,
propensity = pi_test,
type = "posterior",
terms = "tau"
)
t_x_posterior <- colMeans(t_x_posterior)
plot(
tau_0_posterior,
t_x_posterior,
xlab = "tau_0",
ylab = "t(X)",
main = "Posterior of tau_0 vs t(X), averaged over X (Continuous Treatment)"
)