library(stochtree)Bayesian Additive Regression Trees for Supervised Learning
This vignette demonstrates how to sample variants of the BART model (Chipman et al. (2010)), using the bart() function in stochtree. The original BART model is
\[ \begin{aligned} y_i \mid X_i = x_i &\sim \mathcal{N}(f(x_i), \sigma^2)\\ \sigma^2 &\sim \text{IG}\left(\frac{\nu}{2}, \frac{\nu \lambda}{2}\right) \end{aligned} \]
where
\[ f(X) = \sum_{s=1}^m g_s(X) \]
and each \(g_s\) refers to a decision tree function which partitions \(X\) into \(k_s\) mutually exclusive regions (\(\mathcal{A}_s = \mathcal{A}_{s,1} \cup \dots \cup \mathcal{A}_{s,k_s}\)) and assigns a scalar parameter \(\mu_{s,j}\) to each region \(\mathcal{A}_{s,j}\)
\[ g_s(x) = \sum_{j = 1}^{k_s} \mu_{s,j} \mathbb{I}\left(x \in \mathcal{A}_{s,j}\right). \]
The partitions \(\mathcal{A}_s\) are defined by a series of logical split rules \(X_i \leq c\) where \(i\) is a variable index and \(c\) is a numeric cutpoint and these partitions are guided by a uniform prior on variables and cutpoints. The prior on partitions is further specified by a probability of splitting a node
\[ P(\text{split node } \eta) = \alpha (1 + \text{depth}_{\eta})^{-\beta} \]
The prior for each leaf node parameter is
\[ \mu_{s,j} \sim \mathcal{N}\left(0, \sigma^2_{\mu}\right) \]
Together, we refer to this conditional mean model as
\[ f(X) \sim \text{BART}(\alpha, \beta, m) \]
This is the “core” of stochtree’s supervised learning interface, though we support many expanded models including
- linear leaf regression (i.e. each leaf node evaluates a linear regression on basis \(W\) rather than return a constant),
- additive random effects,
- forest-based heteroskedasticity,
- binary / ordinal outcome modeling using the probit and complementary log-log (cloglog) links,
and we offer the ability to sample any of the above models using the MCMC or the Grow-From-Root sampler (He and Hahn (2023)).
Setup
To begin, we load the stochtree and other necessary packages.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from stochtree import BARTModelWe set a seed for reproducibility
random_seed <- 1234
set.seed(random_seed)random_seed = 1234
rng = np.random.default_rng(random_seed)Demo 1: Step Function
Data Generation
We generate data from a simple step function
# Generate the data
n <- 500
p_x <- 10
snr <- 3
X <- matrix(runif(n * p_x), ncol = p_x)
f_XW <- (((0 <= X[, 1]) & (0.25 > X[, 1])) *
(-7.5) +
((0.25 <= X[, 1]) & (0.5 > X[, 1])) * (-2.5) +
((0.5 <= X[, 1]) & (0.75 > X[, 1])) * (2.5) +
((0.75 <= X[, 1]) & (1 > X[, 1])) * (7.5))
noise_sd <- sd(f_XW) / snr
y <- f_XW + rnorm(n, 0, 1) * noise_sd# Generate the data
n = 500
p_x = 10
snr = 3
X = rng.uniform(0, 1, (n, p_x))
f_XW = (
((X[:, 0] >= 0.0) & (X[:, 0] < 0.25)) * (-7.5)
+ ((X[:, 0] >= 0.25) & (X[:, 0] < 0.5)) * (-2.5)
+ ((X[:, 0] >= 0.5) & (X[:, 0] < 0.75)) * (2.5)
+ ((X[:, 0] >= 0.75) & (X[:, 0] < 1.0)) * (7.5)
)
noise_sd = np.std(f_XW) / snr
y = f_XW + rng.normal(0, noise_sd, n)Split the data into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct * n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- as.data.frame(X[test_inds, ])
X_train <- as.data.frame(X[train_inds, ])
y_test <- y[test_inds]
y_train <- y[train_inds]test_set_pct = 0.2
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_set_pct, random_state=random_seed
)Sampling and Analysis
We sample from a BART model of \(y \mid X\) with 10 grow-from-root GFR samples (He and Hahn (2023)) followed by 100 MCMC samples (this is the default in stochtree), run for 4 chains initialized by different GFR iterations.
We also specify \(m = 100\) trees and we let both \(\sigma^2\) and \(\sigma^2_{\mu}\) be updated by Gibbs samplers.
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 100
general_params <- list(
sample_sigma2_global = T,
num_threads = 1,
num_chains = 4,
random_seed = random_seed
)
mean_forest_params <- list(sample_sigma2_leaf = T, num_trees = 100)
bart_model <- stochtree::bart(
X_train = X_train,
y_train = y_train,
X_test = X_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
mean_forest_params = mean_forest_params
)num_gfr = 10
num_burnin = 0
num_mcmc = 100
general_params = {
"sample_sigma2_global": True,
"num_threads": 1,
"num_chains": 4,
"random_seed": random_seed,
}
mean_forest_params = {"sample_sigma2_leaf": True, "num_trees": 100}
bart_model = BARTModel()
bart_model.sample(
X_train=X_train,
y_train=y_train,
X_test=X_test,
num_gfr=num_gfr,
num_burnin=num_burnin,
num_mcmc=num_mcmc,
general_params=general_params,
mean_forest_params=mean_forest_params,
)Plot the mean outcome predictions versus the true outcomes
y_hat_test <- predict(
bart_model,
X = X_test,
terms = "y_hat",
type = "mean"
)
plot(
y_hat_test,
y_test,
xlab = "predicted",
ylab = "actual",
main = "Outcome"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)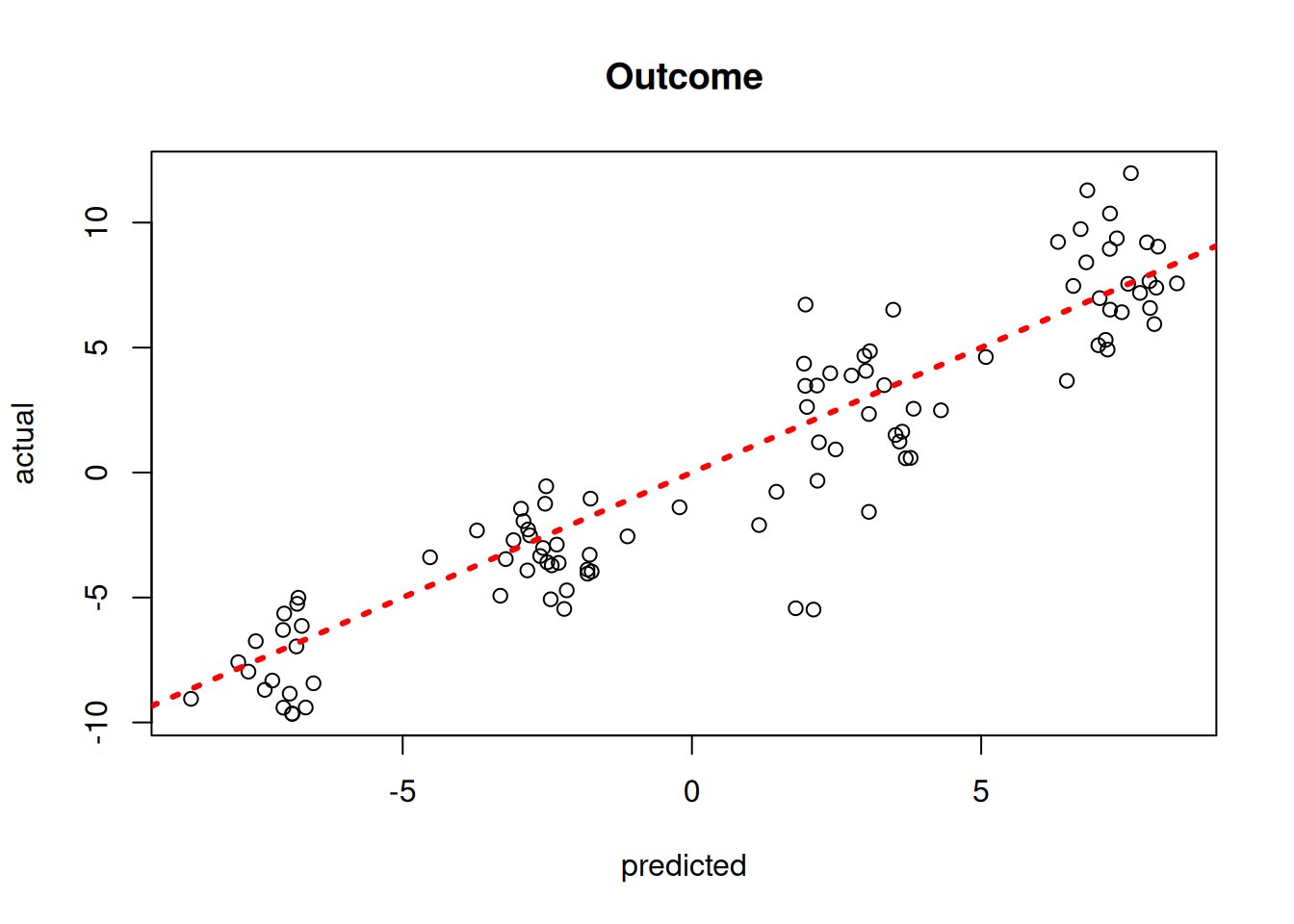
y_hat_test = bart_model.predict(X=X_test, terms="y_hat", type="mean")
lo, hi = min(y_hat_test.min(), y_test.min()), max(y_hat_test.max(), y_test.max())
plt.scatter(y_hat_test, y_test, alpha=0.5)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Outcome")
plt.show()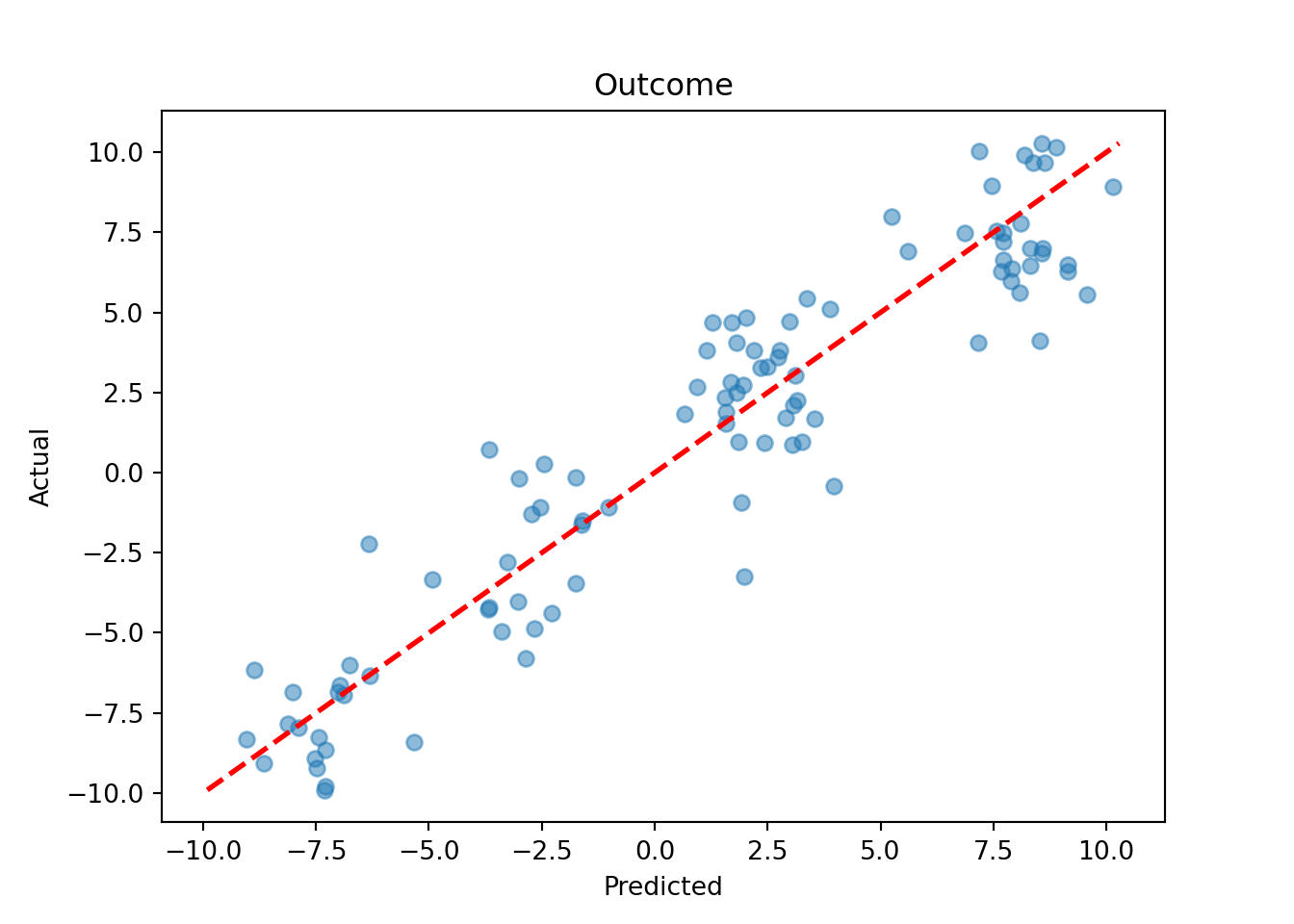
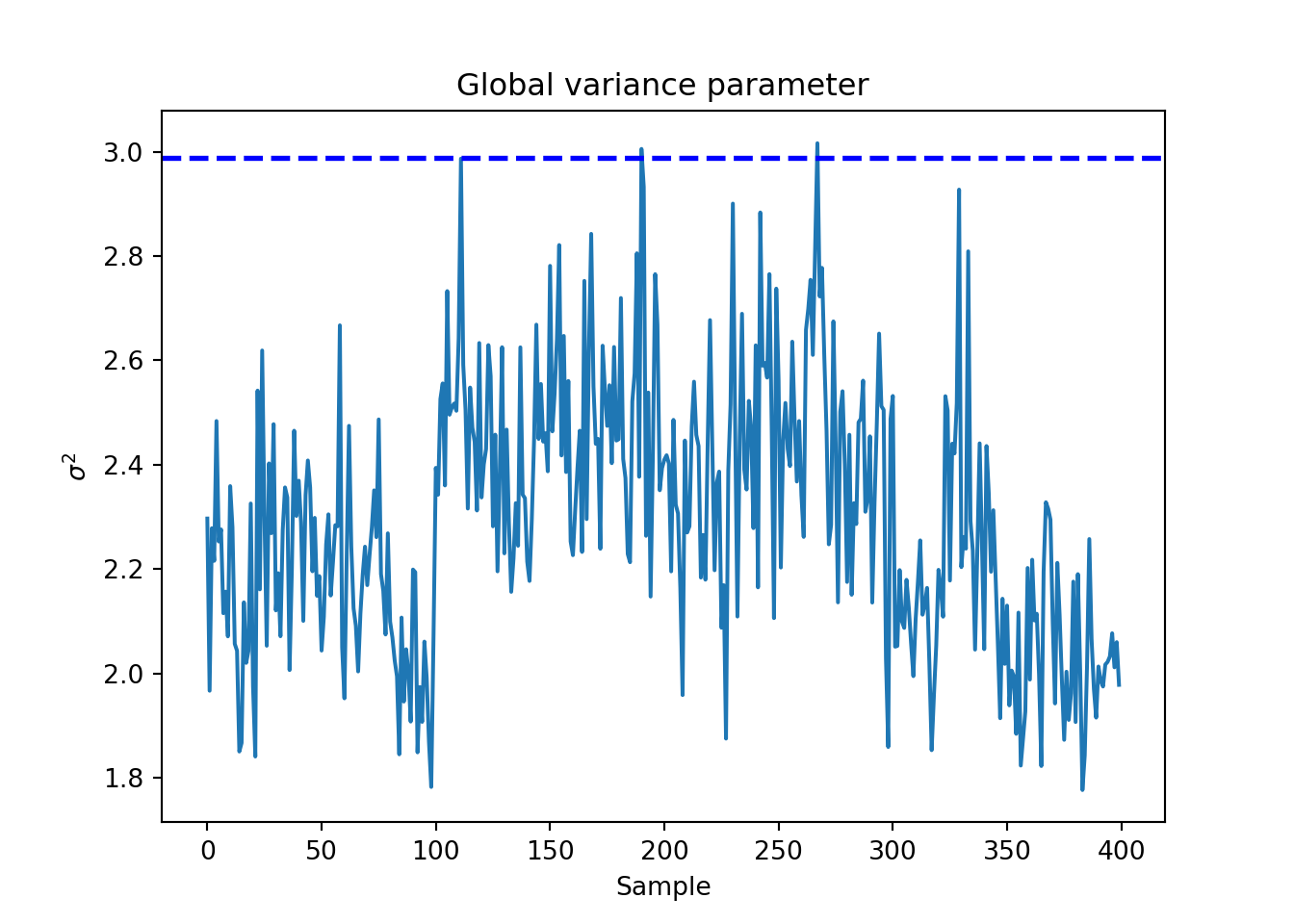
Plot the \(\sigma^2\) traceplot
sigma_observed <- var(y - f_XW)
sigma2_global_samples <- extractParameter(bart_model, "sigma2_global")
plot_bounds <- c(
min(c(sigma2_global_samples, sigma_observed)),
max(c(sigma2_global_samples, sigma_observed))
)
plot(
sigma2_global_samples,
ylim = plot_bounds,
ylab = "sigma^2",
xlab = "Sample",
main = "Global variance parameter"
)
abline(h = sigma_observed, lty = 3, lwd = 3, col = "blue")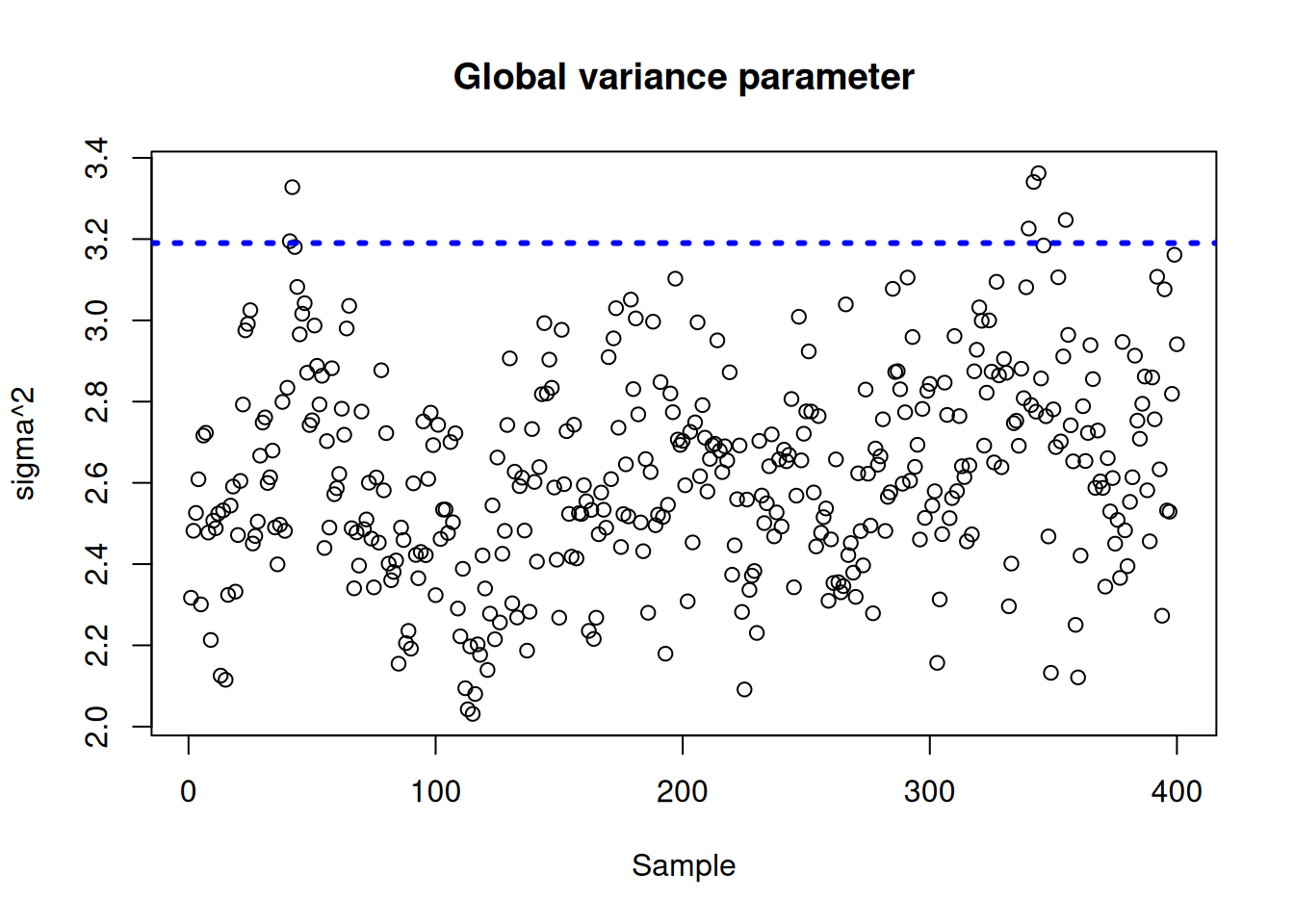
sigma_observed = np.var(y - f_XW)
global_var_samples = bart_model.extract_parameter("sigma2_global")
plt.plot(global_var_samples)
plt.axhline(sigma_observed, color="blue", linestyle="dashed", linewidth=2)
plt.xlabel("Sample")
plt.ylabel(r"$\sigma^2$")
plt.title("Global variance parameter")
plt.show()