library(stochtree)BART with a Forest-based Variance Model
This vignette demonstrates how to configure a “variance forest” in stochtree for modeling conditional variance (see Murray (2021)).
Setup
Load necessary packages
import numpy as np
import matplotlib.pyplot as plt
from stochtree import BARTModelSet a random seed
random_seed = 1234
set.seed(random_seed)random_seed = 1234
rng = np.random.default_rng(random_seed)Demo 1: Variance-Only Simulation (Simple DGP)
Here, we generate data with a constant (zero) mean and a relatively simple covariate-modified variance function.
\[\begin{equation*} \begin{aligned} y &= 0 + \sigma(X) \epsilon\\ \sigma^2(X) &= \begin{cases} 0.5 & X_1 \geq 0 \text{ and } X_1 < 0.25\\ 1 & X_1 \geq 0.25 \text{ and } X_1 < 0.5\\ 2 & X_1 \geq 0.5 \text{ and } X_1 < 0.75\\ 3 & X_1 \geq 0.75 \text{ and } X_1 < 1\\ \end{cases}\\ X_1,\dots,X_p &\sim \text{U}\left(0,1\right)\\ \epsilon &\sim \mathcal{N}\left(0,1\right) \end{aligned} \end{equation*}\]
Simulation
Generate data from the DGP above
n <- 1000
p_x <- 10
X <- matrix(runif(n * p_x), ncol = p_x)
f_XW <- 0
s_XW <- (((0 <= X[, 1]) & (0.25 > X[, 1])) *
(0.5) +
((0.25 <= X[, 1]) & (0.5 > X[, 1])) * (1) +
((0.5 <= X[, 1]) & (0.75 > X[, 1])) * (2) +
((0.75 <= X[, 1]) & (1 > X[, 1])) * (3))
y <- f_XW + rnorm(n, 0, 1) * s_XWn, p_x = 1000, 10
X = rng.uniform(size=(n, p_x))
s_XW = (
((X[:, 0] >= 0) & (X[:, 0] < 0.25)) * 0.5
+ ((X[:, 0] >= 0.25) & (X[:, 0] < 0.5)) * 1.0
+ ((X[:, 0] >= 0.5) & (X[:, 0] < 0.75)) * 2.0
+ ((X[:, 0] >= 0.75) & (X[:, 0] < 1.0)) * 3.0
)
y = rng.normal(size=n) * s_XWSplit into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct * n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- as.data.frame(X[test_inds, ])
X_train <- as.data.frame(X[train_inds, ])
y_test <- y[test_inds]
y_train <- y[train_inds]
s_x_test <- s_XW[test_inds]
s_x_train <- s_XW[train_inds]test_set_pct = 0.2
n_test = round(test_set_pct * n)
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test, X_train = X[test_inds], X[train_inds]
y_test, y_train = y[test_inds], y[train_inds]
s_x_test, s_x_train = s_XW[test_inds], s_XW[train_inds]Sampling and Analysis
We sample four chains of the \(\sigma^2(X)\) forest using “warm-start” initialization (He and Hahn (2023)).
We use fewer trees for the variance forest than typically used for mean forests, and we disable sampling a global error scale and omit the mean forest by setting num_trees = 0 in its parameter list.
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 100
num_trees <- 20
num_samples <- num_gfr + num_burnin + num_mcmc
general_params <- list(
sample_sigma2_global = F,
num_chains = 4,
num_threads = 1,
random_seed = random_seed
)
mean_forest_params <- list(sample_sigma2_leaf = F, num_trees = 0)
variance_forest_params <- list(num_trees = num_trees)
bart_model <- stochtree::bart(
X_train = X_train,
y_train = y_train,
X_test = X_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
mean_forest_params = mean_forest_params,
variance_forest_params = variance_forest_params
)num_gfr = 10
num_burnin = 0
num_mcmc = 100
num_trees = 20
bart_model = BARTModel()
bart_model.sample(
X_train=X_train,
y_train=y_train,
X_test=X_test,
num_gfr=num_gfr,
num_burnin=num_burnin,
num_mcmc=num_mcmc,
general_params={
"sample_sigma2_global": False,
"num_threads": 1,
"num_chains": 4,
"random_seed": random_seed,
},
mean_forest_params={"sample_sigma2_leaf": False, "num_trees": 0},
variance_forest_params={"num_trees": num_trees},
)We inspect the model by plotting the true variance function against its forest-based predictions
sigma2_x_hat_test <- predict(
bart_model,
X = X_test,
terms = "variance_forest",
type = "mean"
)
plot(
sigma2_x_hat_test,
s_x_test^2,
pch = 16,
cex = 0.75,
xlab = "Predicted",
ylab = "Actual",
main = "Variance function"
)
abline(0, 1, col = "red", lty = 2, lwd = 2.5)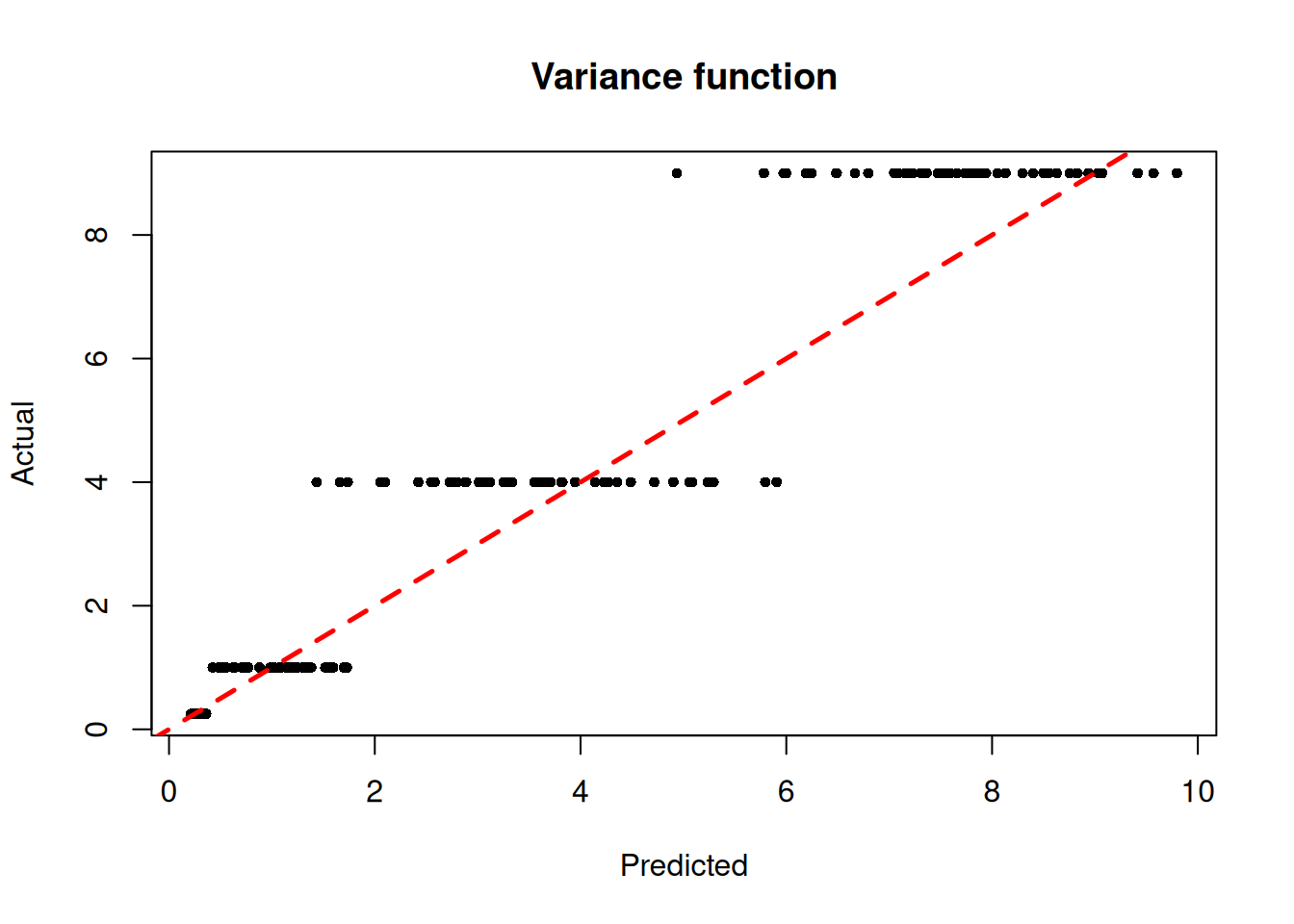
sigma2_x_hat_test = bart_model.predict(X=X_test, terms="variance_forest", type="mean")
lo, hi = (
min(sigma2_x_hat_test.min(), (s_x_test**2).min()),
max(sigma2_x_hat_test.max(), (s_x_test**2).max()),
)
plt.scatter(sigma2_x_hat_test, s_x_test**2, s=10, alpha=0.6)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Variance function")
plt.show()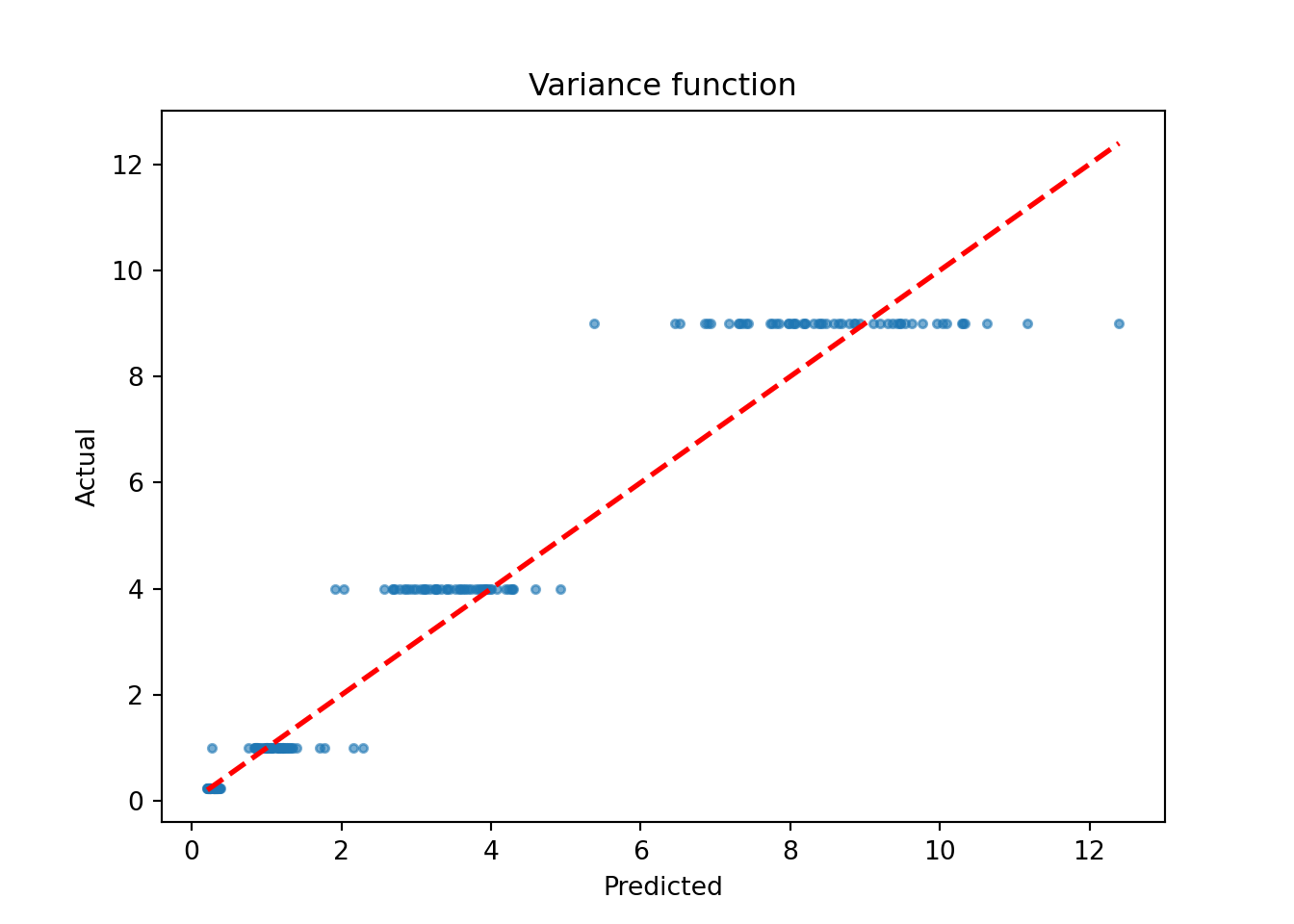
Demo 2: Variance-Only Simulation (Complex DGP)
Here, we generate data with a constant (zero) mean and a more complex covariate-modified variance function.
\[\begin{equation*} \begin{aligned} y &= 0 + \sigma(X) \epsilon\\ \sigma^2(X) &= \begin{cases} 0.25X_3^2 & X_1 \geq 0 \text{ and } X_1 < 0.25\\ 1X_3^2 & X_1 \geq 0.25 \text{ and } X_1 < 0.5\\ 4X_3^2 & X_1 \geq 0.5 \text{ and } X_1 < 0.75\\ 9X_3^2 & X_1 \geq 0.75 \text{ and } X_1 < 1\\ \end{cases}\\ X_1,\dots,X_p &\sim \text{U}\left(0,1\right)\\ \epsilon &\sim \mathcal{N}\left(0,1\right) \end{aligned} \end{equation*}\]
Simulation
We generate data from the DGP above
n <- 1000
p_x <- 10
X <- matrix(runif(n*p_x), ncol = p_x)
f_XW <- 0
s_XW <- (
((0 <= X[,1]) & (0.25 > X[,1])) * (0.5*X[,3]) +
((0.25 <= X[,1]) & (0.5 > X[,1])) * (1*X[,3]) +
((0.5 <= X[,1]) & (0.75 > X[,1])) * (2*X[,3]) +
((0.75 <= X[,1]) & (1 > X[,1])) * (3*X[,3])
)
y <- f_XW + rnorm(n, 0, 1)*s_XWn, p_x = 1000, 10
X = rng.uniform(size=(n, p_x))
# R's X[,3] = Python's X[:,2]
s_XW = (
((X[:, 0] >= 0) & (X[:, 0] < 0.25)) * (0.5 * X[:, 2]) +
((X[:, 0] >= 0.25) & (X[:, 0] < 0.5)) * (1.0 * X[:, 2]) +
((X[:, 0] >= 0.5) & (X[:, 0] < 0.75)) * (2.0 * X[:, 2]) +
((X[:, 0] >= 0.75) & (X[:, 0] < 1.0)) * (3.0 * X[:, 2])
)
y = rng.normal(size=n) * s_XWAnd split the data into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct*n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- as.data.frame(X[test_inds,])
X_train <- as.data.frame(X[train_inds,])
y_test <- y[test_inds]
y_train <- y[train_inds]
s_x_test <- s_XW[test_inds]
s_x_train <- s_XW[train_inds]test_set_pct = 0.2
n_test = round(test_set_pct * n)
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test, X_train = X[test_inds], X[train_inds]
y_test, y_train = y[test_inds], y[train_inds]
s_x_test, s_x_train = s_XW[test_inds], s_XW[train_inds]Sampling and Analysis
We sample four chains of the \(\sigma^2(X)\) forest using “warm-start” initialization (He and Hahn (2023)).
We use fewer trees for the variance forest than typically used for mean forests, and we disable sampling a global error scale and omit the mean forest by setting num_trees = 0 in its parameter list.
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 100
num_trees <- 20
num_samples <- num_gfr + num_burnin + num_mcmc
general_params <- list(
sample_sigma2_global = F,
num_chains = 4,
num_threads = 1,
random_seed = random_seed
)
mean_forest_params <- list(sample_sigma2_leaf = F, num_trees = 0)
variance_forest_params <- list(num_trees = num_trees)
bart_model <- stochtree::bart(
X_train = X_train,
y_train = y_train,
X_test = X_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
mean_forest_params = mean_forest_params,
variance_forest_params = variance_forest_params
)num_gfr = 10
num_burnin = 0
num_mcmc = 100
num_trees = 20
bart_model = BARTModel()
bart_model.sample(
X_train=X_train,
y_train=y_train,
X_test=X_test,
num_gfr=num_gfr,
num_burnin=num_burnin,
num_mcmc=num_mcmc,
general_params={
"sample_sigma2_global": False,
"num_threads": 1,
"num_chains": 4,
"random_seed": random_seed,
},
mean_forest_params={"sample_sigma2_leaf": False, "num_trees": 0},
variance_forest_params={"num_trees": num_trees},
)We inspect the model by plotting the true variance function against its forest-based predictions
sigma2_x_hat_test <- predict(
bart_model,
X = X_test,
terms = "variance_forest",
type = "mean"
)
plot(
sigma2_x_hat_test,
s_x_test^2,
pch = 16,
cex = 0.75,
xlab = "Predicted",
ylab = "Actual",
main = "Variance function"
)
abline(0, 1, col = "red", lty = 2, lwd = 2.5)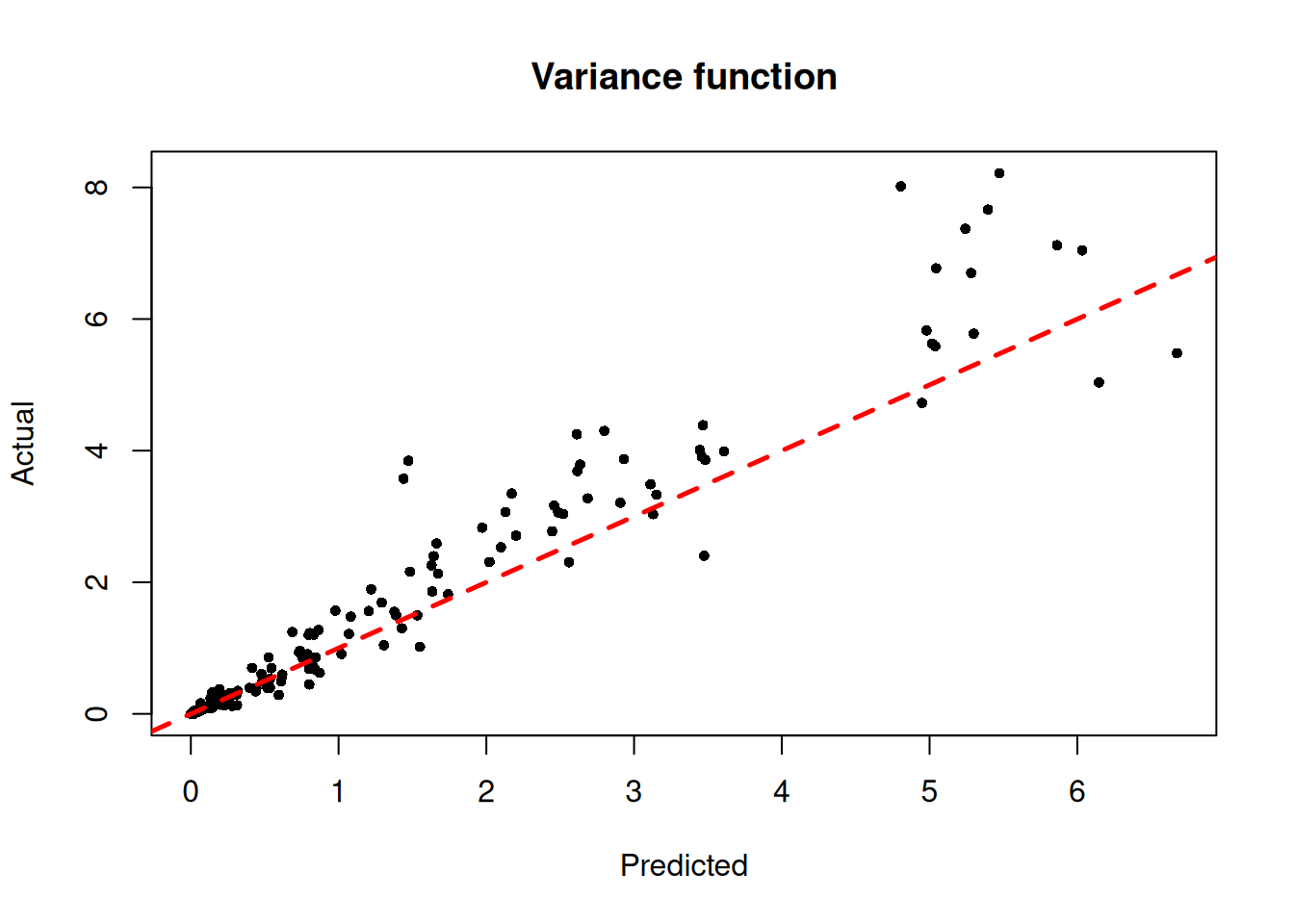
sigma2_x_hat_test = bart_model.predict(X=X_test, terms="variance_forest", type="mean")
lo, hi = (
min(sigma2_x_hat_test.min(), (s_x_test**2).min()),
max(sigma2_x_hat_test.max(), (s_x_test**2).max()),
)
plt.scatter(sigma2_x_hat_test, s_x_test**2, s=10, alpha=0.6)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Variance function")
plt.show()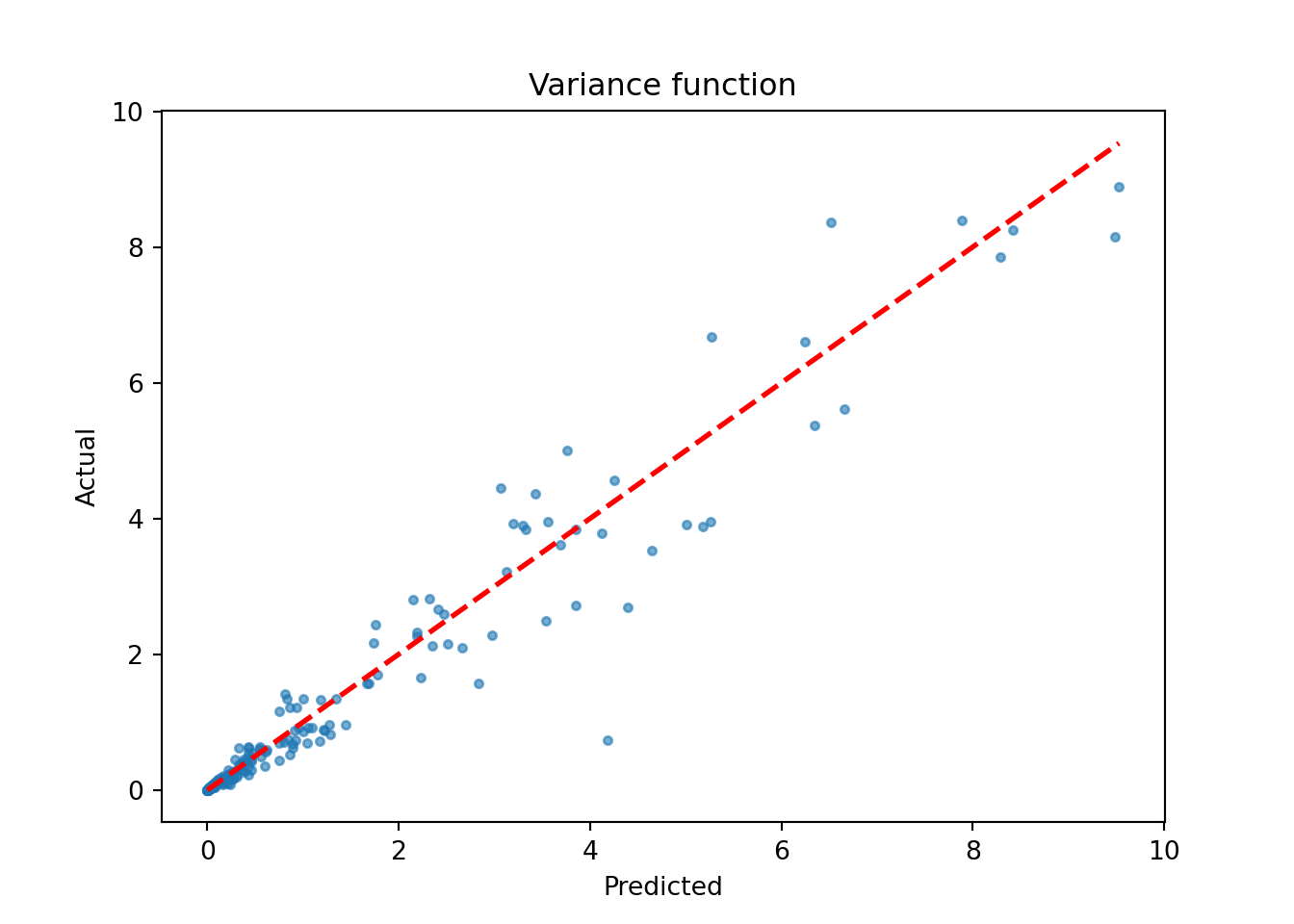
Demo 3: Mean and Variance Function Simulation
Here, we generate data with (relatively simple) covariate-modified mean and variance functions.
\[\begin{equation*} \begin{aligned} y &= f(X) + \sigma(X) \epsilon\\ f(X) &= \begin{cases} -6 & X_2 \geq 0 \text{ and } X_2 < 0.25\\ -2 & X_2 \geq 0.25 \text{ and } X_2 < 0.5\\ 2 & X_2 \geq 0.5 \text{ and } X_2 < 0.75\\ 6 & X_2 \geq 0.75 \text{ and } X_2 < 1\\ \end{cases}\\ \sigma^2(X) &= \begin{cases} 0.25 & X_1 \geq 0 \text{ and } X_1 < 0.25\\ 1 & X_1 \geq 0.25 \text{ and } X_1 < 0.5\\ 4 & X_1 \geq 0.5 \text{ and } X_1 < 0.75\\ 9 & X_1 \geq 0.75 \text{ and } X_1 < 1\\ \end{cases}\\ X_1,\dots,X_p &\sim \text{U}\left(0,1\right)\\ \epsilon &\sim \mathcal{N}\left(0,1\right) \end{aligned} \end{equation*}\]
Simulation
Generate data from the DGP above
n <- 1000
p_x <- 10
X <- matrix(runif(n*p_x), ncol = p_x)
f_XW <- (
((0 <= X[,2]) & (0.25 > X[,2])) * (-6) +
((0.25 <= X[,2]) & (0.5 > X[,2])) * (-2) +
((0.5 <= X[,2]) & (0.75 > X[,2])) * (2) +
((0.75 <= X[,2]) & (1 > X[,2])) * (6)
)
s_XW <- (
((0 <= X[,1]) & (0.25 > X[,1])) * (0.5) +
((0.25 <= X[,1]) & (0.5 > X[,1])) * (1) +
((0.5 <= X[,1]) & (0.75 > X[,1])) * (2) +
((0.75 <= X[,1]) & (1 > X[,1])) * (3)
)
y <- f_XW + rnorm(n, 0, 1)*s_XWn, p_x = 1000, 10
X = rng.uniform(size=(n, p_x))
f_XW = (
((X[:, 1] >= 0) & (X[:, 1] < 0.25)) * (-6) +
((X[:, 1] >= 0.25) & (X[:, 1] < 0.5)) * (-2) +
((X[:, 1] >= 0.5) & (X[:, 1] < 0.75)) * (2) +
((X[:, 1] >= 0.75) & (X[:, 1] < 1.0)) * (6)
)
s_XW = (
((X[:, 0] >= 0) & (X[:, 0] < 0.25)) * 0.5 +
((X[:, 0] >= 0.25) & (X[:, 0] < 0.5)) * 1.0 +
((X[:, 0] >= 0.5) & (X[:, 0] < 0.75)) * 2.0 +
((X[:, 0] >= 0.75) & (X[:, 0] < 1.0)) * 3.0
)
y = f_XW + rng.normal(size=n) * s_XWSplit the data into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct*n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- as.data.frame(X[test_inds,])
X_train <- as.data.frame(X[train_inds,])
y_test <- y[test_inds]
y_train <- y[train_inds]
f_x_test <- f_XW[test_inds]
s_x_test <- s_XW[test_inds]test_set_pct = 0.2
n_test = round(test_set_pct * n)
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test, X_train = X[test_inds], X[train_inds]
y_test, y_train = y[test_inds], y[train_inds]
f_x_test = f_XW[test_inds]
s_x_test = s_XW[test_inds]Sampling and Analysis
As above, we sample four chains of the \(\sigma^2(X)\) forest using “warm-start” initialization (He and Hahn (2023)), except we do not omit the mean forest by setting num_trees = 0.
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 100
general_params <- list(
sample_sigma2_global = F,
num_threads = 1,
num_chains = 4,
random_seed = random_seed
)
mean_forest_params <- list(
sample_sigma2_leaf = F,
num_trees = 50,
alpha = 0.95,
beta = 2,
min_samples_leaf = 5
)
variance_forest_params <- list(
num_trees = 50,
alpha = 0.95,
beta = 1.25,
min_samples_leaf = 5
)
bart_model <- stochtree::bart(
X_train = X_train,
y_train = y_train,
X_test = X_test,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params,
mean_forest_params = mean_forest_params,
variance_forest_params = variance_forest_params
)bart_model = BARTModel()
bart_model.sample(
X_train=X_train,
y_train=y_train,
X_test=X_test,
num_gfr=10,
num_burnin=0,
num_mcmc=100,
general_params={
"sample_sigma2_global": False,
"num_threads": 1,
"num_chains": 4,
"random_seed": random_seed,
},
mean_forest_params={
"sample_sigma2_leaf": False,
"num_trees": 50,
"alpha": 0.95,
"beta": 2,
"min_samples_leaf": 5,
},
variance_forest_params={
"num_trees": 50,
"alpha": 0.95,
"beta": 1.25,
"min_samples_leaf": 5,
},
)We inspect the model by plotting the true variance function against the variance forest predictions
sigma2_x_hat_test <- predict(
bart_model,
X = X_test,
terms = "variance_forest",
type = "mean"
)
plot(
sigma2_x_hat_test,
s_x_test^2,
pch = 16,
cex = 0.75,
xlab = "Predicted",
ylab = "Actual",
main = "Variance function"
)
abline(0, 1, col = "red", lty = 2, lwd = 2.5)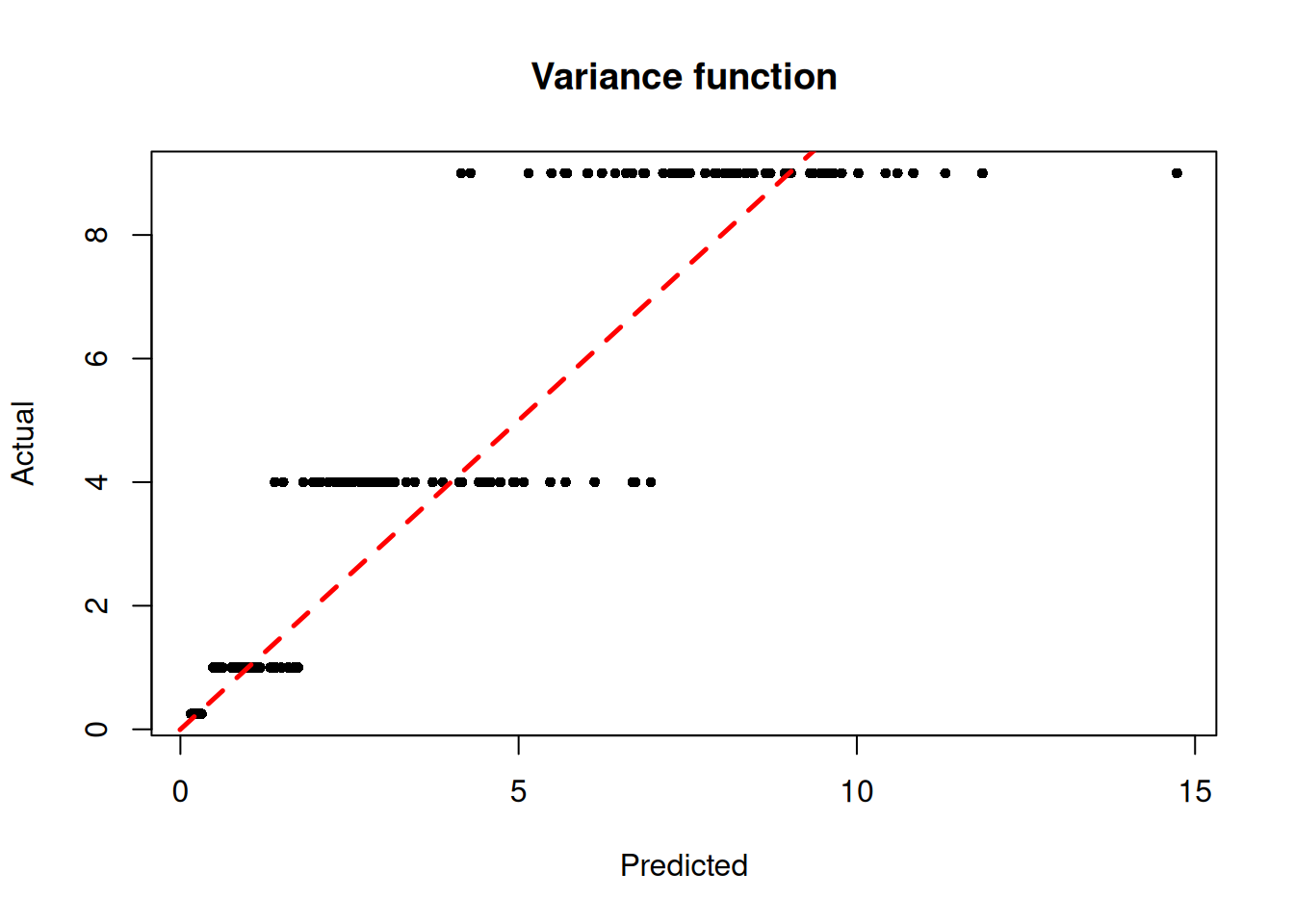
sigma2_x_hat_test = bart_model.predict(X=X_test, terms="variance_forest", type="mean")
lo, hi = (
min(sigma2_x_hat_test.min(), (s_x_test**2).min()),
max(sigma2_x_hat_test.max(), (s_x_test**2).max()),
)
plt.scatter(sigma2_x_hat_test, s_x_test**2, s=10, alpha=0.6)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Variance function")
plt.show()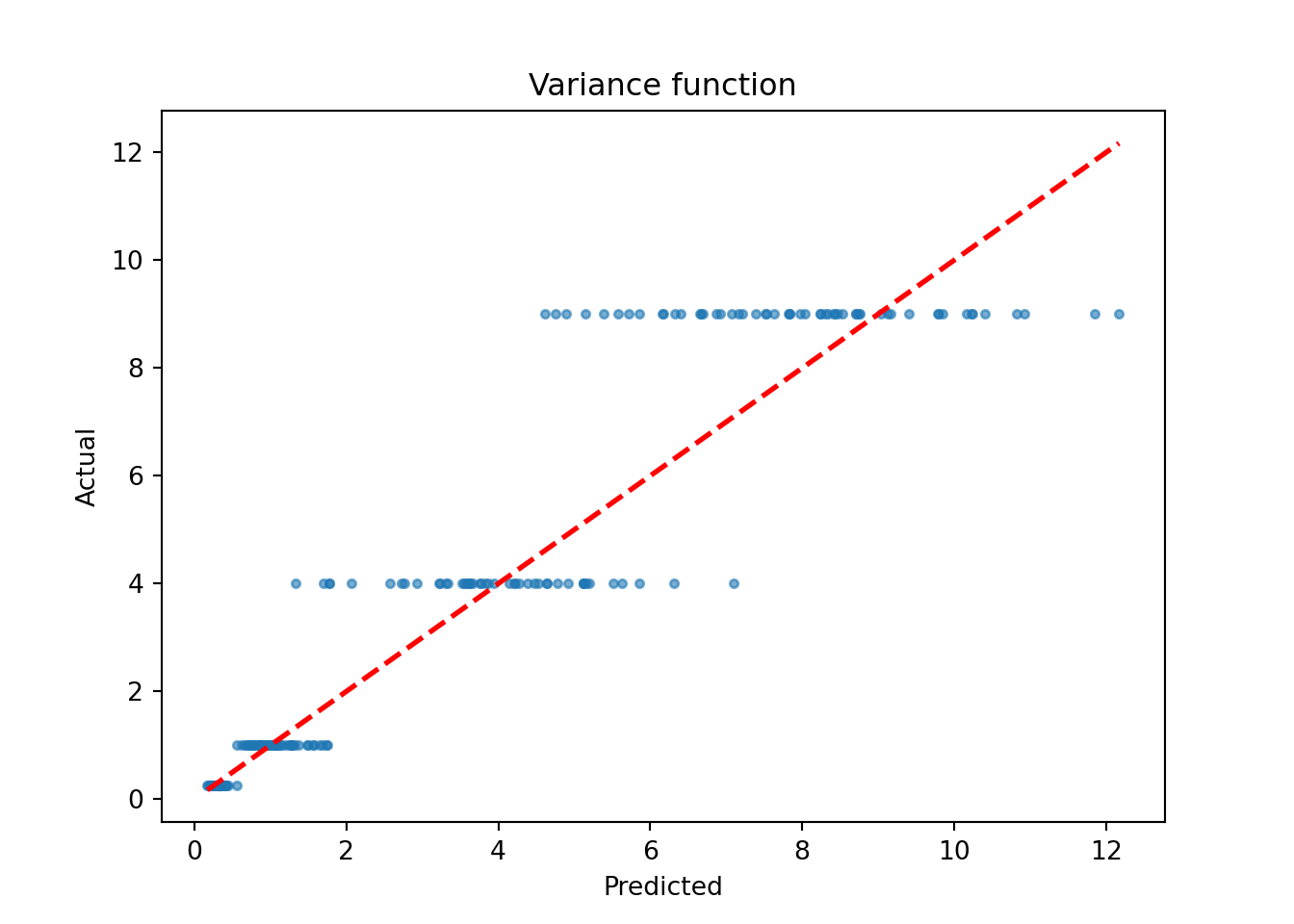
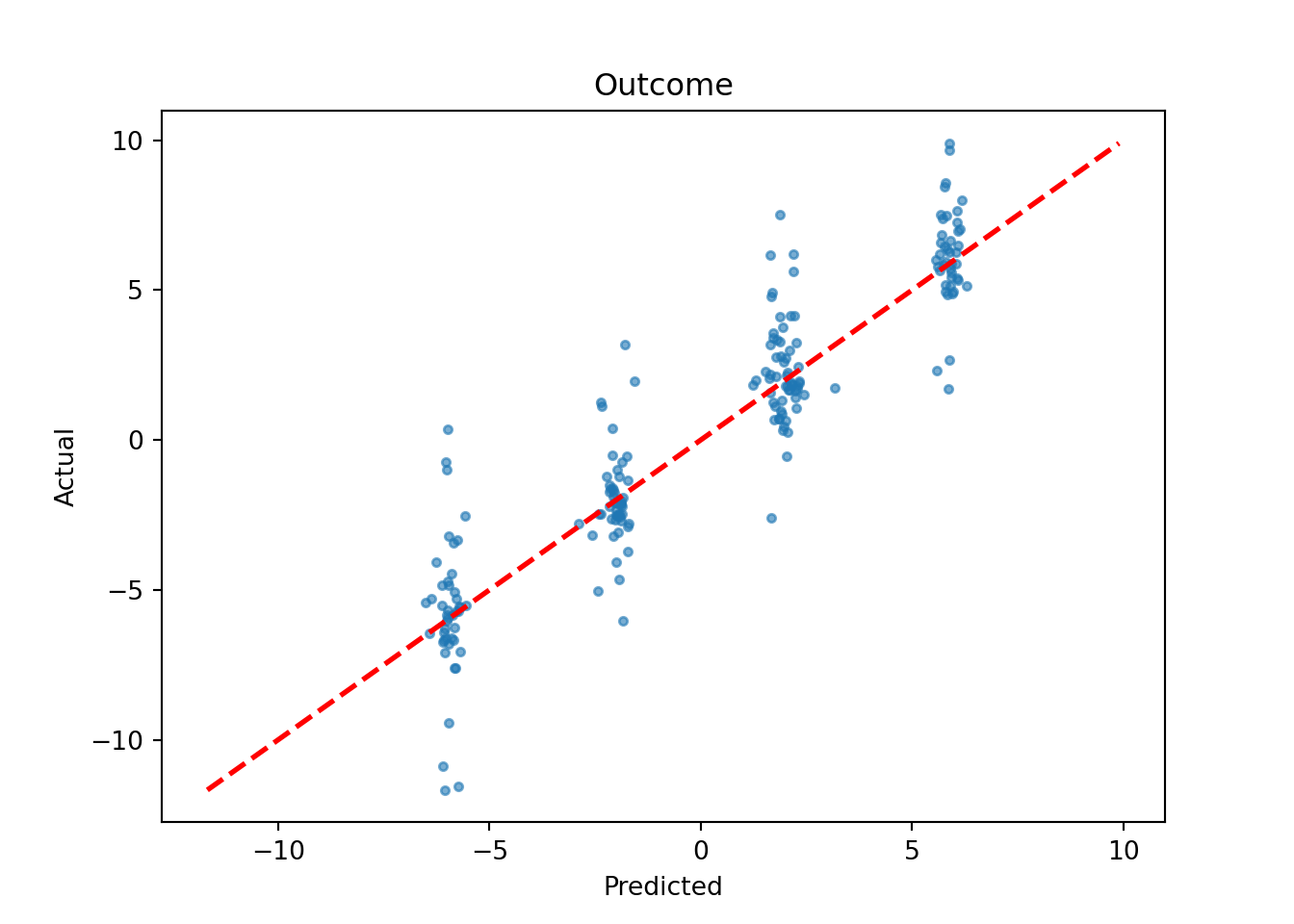
We also plot the true outcome against mean forest predictions
y_hat_test <- predict(
bart_model,
X = X_test,
terms = "y_hat",
type = "mean"
)
plot(
y_hat_test,
y_test,
pch = 16,
cex = 0.75,
xlab = "Predicted",
ylab = "Actual",
main = "Outcome"
)
abline(0, 1, col = "red", lty = 2, lwd = 2.5)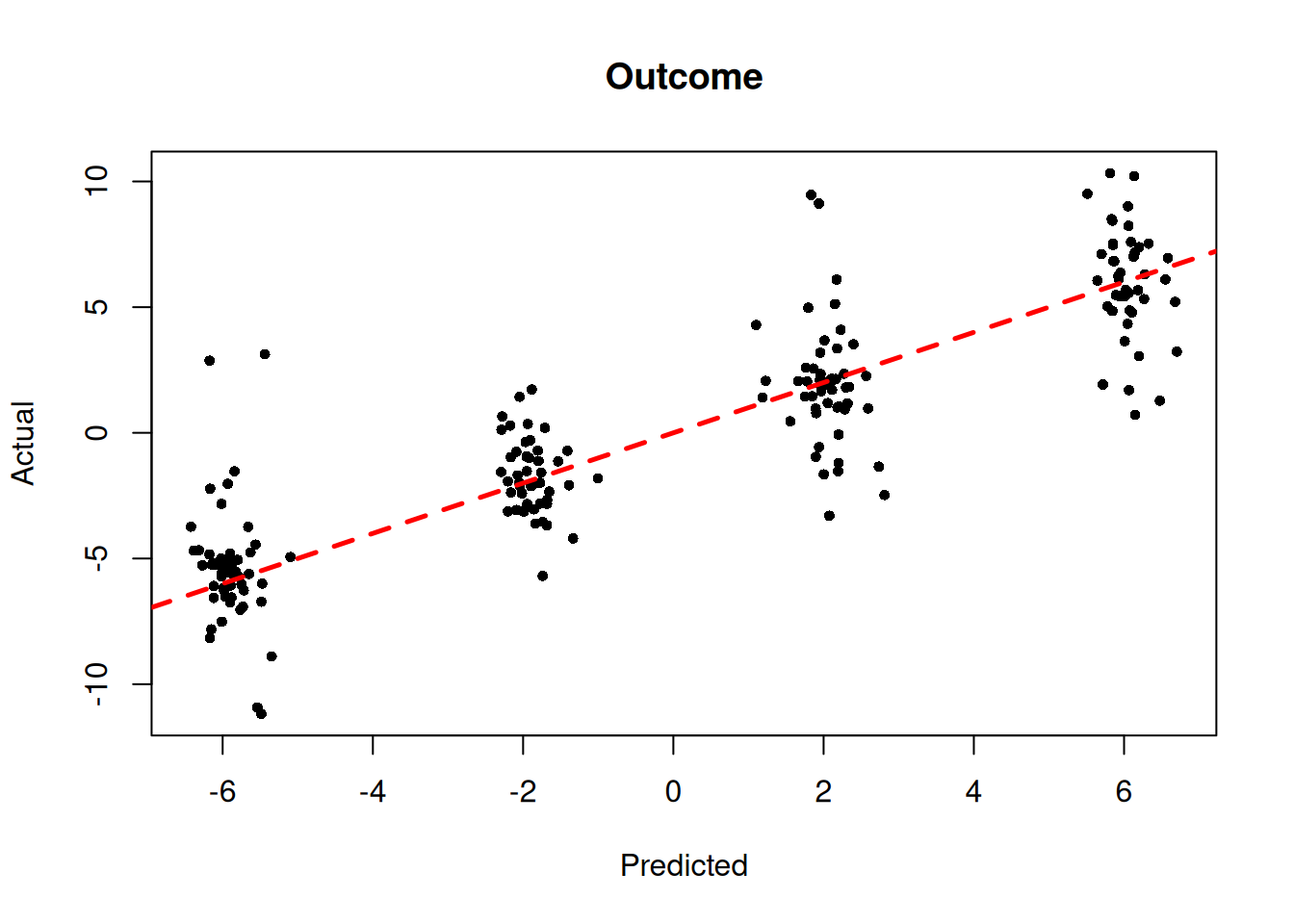
y_hat_test = bart_model.predict(X=X_test, terms="y_hat", type="mean")
lo, hi = (
min(y_hat_test.min(), y_test.min()),
max(y_hat_test.max(), y_test.max()),
)
plt.scatter(y_hat_test, y_test, s=10, alpha=0.6)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Outcome")
plt.show()