library(stochtree)Bayesian Causal Forests for Treatment Effect Estimation
This vignette demonstrates how to use the bcf() function for causal inference (Hahn et al. (2020)). BCF models the conditional average treatment effect (CATE) by fitting two separate tree ensembles
\[ Y_i = \mu(X_i) + \tau(X_i) Z_i + \epsilon_i, \quad \epsilon_i \sim \mathcal{N}(0, \sigma^2) \]
where \(\mu(\cdot)\) is a prognostic forest and \(\tau(\cdot)\) is a treatment effect forest. The estimated propensity score \(\hat{\pi}(X_i)\) is included as a covariate in \(\mu(\cdot)\) to reduce confounding bias.
Setup
Load necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import norm
from stochtree import BCFModelSet a seed for reproducibility
random_seed <- 1234
set.seed(random_seed)random_seed = 1234
rng = np.random.default_rng(random_seed)We also define several simple functions that configure the data generating processes used in this vignette
g <- function(x) {
ifelse(x[, 5] == 1, 2, ifelse(x[, 5] == 2, -1, -4))
}
mu1 <- function(x) {
1 + g(x) + x[, 1] * x[, 3]
}
mu2 <- function(x) {
1 + g(x) + 6 * abs(x[, 3] - 1)
}
tau1 <- function(x) {
rep(3, nrow(x))
}
tau2 <- function(x) {
1 + 2 * x[, 2] * x[, 4]
}def g(x):
return np.where(x[:, 4] == 1, 2, np.where(x[:, 4] == 2, -1, -4))
def mu1(x):
return 1 + g(x) + x[:, 0] * x[:, 2]
def mu2(x):
return 1 + g(x) + 6 * np.abs(x[:, 2] - 1)
def tau1(x):
return np.full(x.shape[0], 3.0)
def tau2(x):
return 1 + 2 * x[:, 1] * x[:, 3]Binary Treatment
Demo 1: Linear Outcome Model, Heterogeneous Treatment Effect
We consider the following data generating process from Hahn et al. (2020):
\[\begin{equation*} \begin{aligned} y &= \mu(X) + \tau(X) Z + \epsilon\\ \epsilon &\sim N\left(0,\sigma^2\right)\\ \mu(X) &= 1 + g(X) + 6 X_1 X_3\\ \tau(X) &= 1 + 2 X_2 X_4\\ g(X) &= \mathbb{I}(X_5=1) \times 2 - \mathbb{I}(X_5=2) \times 1 - \mathbb{I}(X_5=3) \times 4\\ s_{\mu} &= \sqrt{\mathbb{V}(\mu(X))}\\ \pi(X) &= 0.8 \phi\left(\frac{3\mu(X)}{s_{\mu}}\right) - \frac{X_1}{2} + \frac{2U+1}{20}\\ X_1,X_2,X_3 &\sim N\left(0,1\right)\\ X_4 &\sim \text{Bernoulli}(1/2)\\ X_5 &\sim \text{Categorical}(1/3,1/3,1/3)\\ U &\sim \text{Uniform}\left(0,1\right)\\ Z &\sim \text{Bernoulli}\left(\pi(X)\right) \end{aligned} \end{equation*}\]
Simulation
We generate data from the DGP defined above
n <- 1000
snr <- 3
x1 <- rnorm(n)
x2 <- rnorm(n)
x3 <- rnorm(n)
x4 <- as.numeric(rbinom(n, 1, 0.5))
x5 <- as.numeric(sample(1:3, n, replace = TRUE))
X <- cbind(x1, x2, x3, x4, x5)
p <- ncol(X)
mu_x <- mu1(X)
tau_x <- tau2(X)
pi_x <- 0.8 * pnorm((3 * mu_x / sd(mu_x)) - 0.5 * X[, 1]) + 0.05 + runif(n) / 10
Z <- rbinom(n, 1, pi_x)
E_XZ <- mu_x + Z * tau_x
y <- E_XZ + rnorm(n, 0, 1) * (sd(E_XZ) / snr)
X <- as.data.frame(X)
X$x4 <- factor(X$x4, ordered = TRUE)
X$x5 <- factor(X$x5, ordered = TRUE)n = 1000
snr = 3
x1 = rng.normal(size=n)
x2 = rng.normal(size=n)
x3 = rng.normal(size=n)
x4 = rng.binomial(1, 0.5, n).astype(float)
x5 = rng.choice([1, 2, 3], size=n).astype(float)
X = np.column_stack([x1, x2, x3, x4, x5])
mu_x = mu1(X)
tau_x = tau2(X)
pi_x = (
0.8 * norm.cdf((3 * mu_x / np.std(mu_x)) - 0.5 * X[:, 0])
+ 0.05
+ rng.uniform(size=n) / 10
)
Z = rng.binomial(1, pi_x, n).astype(float)
E_XZ = mu_x + Z * tau_x
y = E_XZ + rng.normal(size=n) * (np.std(E_XZ) / snr)
X_df = pd.DataFrame({"x1": x1, "x2": x2, "x3": x3, "x4": x4, "x5": x5})
X_df["x4"] = pd.Categorical(X_df["x4"].astype(int), categories=[0, 1], ordered=True)
X_df["x5"] = pd.Categorical(X_df["x5"].astype(int), categories=[1, 2, 3], ordered=True)Split data into test and train sets
test_set_pct <- 0.2
n_test <- round(test_set_pct * n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- X[test_inds, ]
X_train <- X[train_inds, ]
pi_test <- pi_x[test_inds]
pi_train <- pi_x[train_inds]
Z_test <- Z[test_inds]
Z_train <- Z[train_inds]
y_test <- y[test_inds]
y_train <- y[train_inds]
mu_test <- mu_x[test_inds]
mu_train <- mu_x[train_inds]
tau_test <- tau_x[test_inds]
tau_train <- tau_x[train_inds]test_set_pct = 0.2
n_test = round(test_set_pct * n)
n_train = n - n_test
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test = X_df.iloc[test_inds]
X_train = X_df.iloc[train_inds]
pi_test, pi_train = pi_x[test_inds], pi_x[train_inds]
Z_test, Z_train = Z[test_inds], Z[train_inds]
y_test, y_train = y[test_inds], y[train_inds]
mu_test, mu_train = mu_x[test_inds], mu_x[train_inds]
tau_test, tau_train = tau_x[test_inds], tau_x[train_inds]Sampling and Analysis
We simulate from a BCF model initialized by “warm-start” samples fit with the grow-from-root algorithm (He and Hahn (2023), Krantsevich et al. (2023)). This is the default in stochtree.
general_params <- list(
num_threads=1,
num_chains=4,
random_seed=random_seed
)
bcf_model <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
num_gfr = 10,
num_burnin = 1000,
num_mcmc = 100,
propensity_test = pi_test,
general_params = general_params
)general_params = {"num_threads": 1, "num_chains": 4, "random_seed": random_seed}
bcf_model = BCFModel()
bcf_model.sample(
X_train=X_train,
Z_train=Z_train,
y_train=y_train,
propensity_train=pi_train,
X_test=X_test,
Z_test=Z_test,
num_gfr=10,
num_burnin=1000,
num_mcmc=100,
propensity_test=pi_test,
general_params=general_params,
)Plot the true versus estimated prognostic function
mu_hat_test <- predict(
bcf_model,
X = X_test,
Z = Z_test,
propensity = pi_test,
terms = "prognostic_function"
)
plot(
rowMeans(mu_hat_test),
mu_test,
xlab = "predicted",
ylab = "actual",
main = "Prognostic function"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)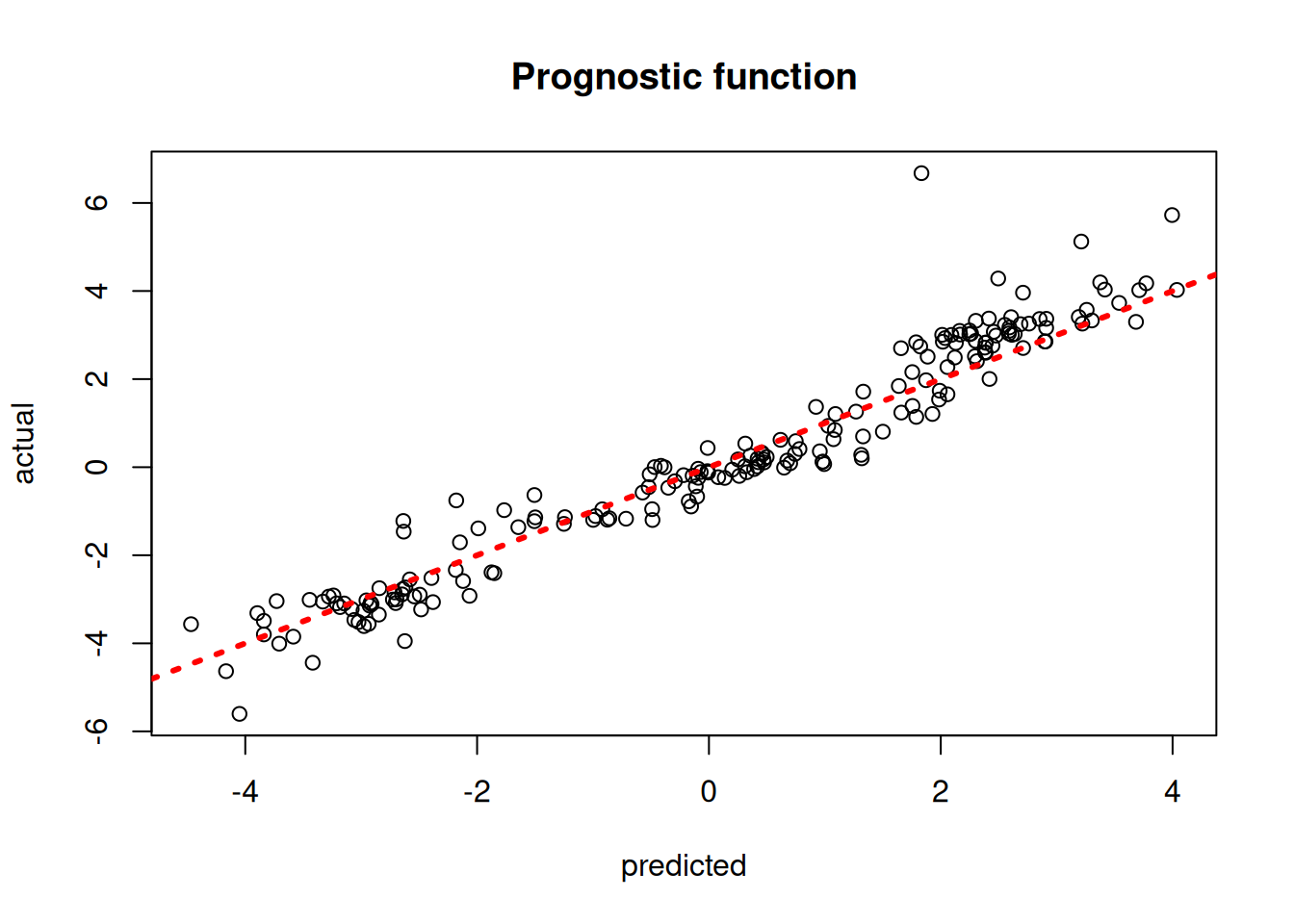
mu_hat_test = bcf_model.predict(
X=X_test, Z=Z_test, propensity=pi_test, terms="prognostic_function"
)
mu_pred = mu_hat_test.mean(axis=1)
lo, hi = min(mu_pred.min(), mu_test.min()), max(mu_pred.max(), mu_test.max())
plt.scatter(mu_pred, mu_test, alpha=0.5)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Prognostic function")
plt.show()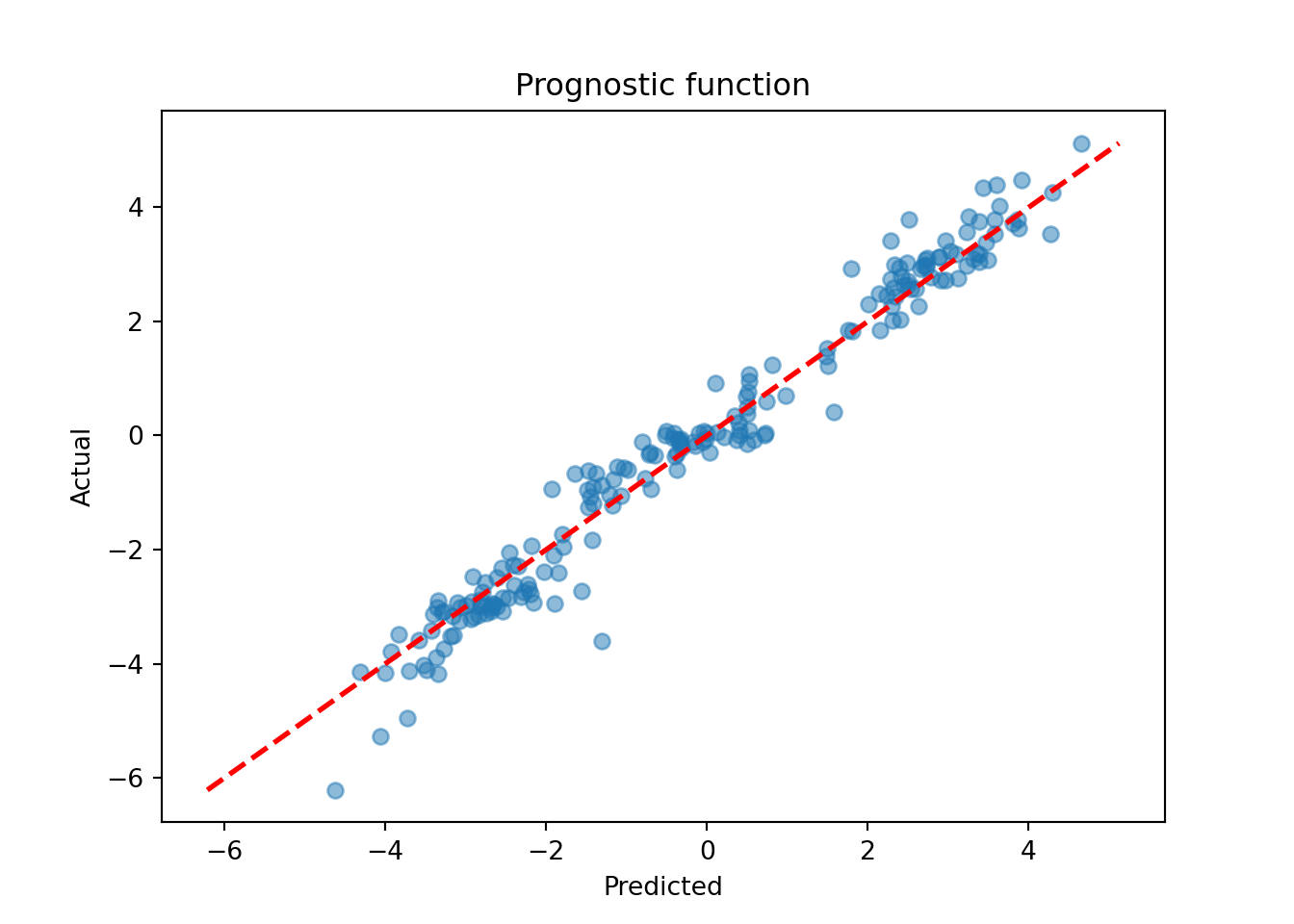
Plot the true versus estimated CATE function
tau_hat_test <- predict(
bcf_model,
X = X_test,
Z = Z_test,
propensity = pi_test,
terms = "cate"
)
plot(
rowMeans(tau_hat_test),
tau_test,
xlab = "predicted",
ylab = "actual",
main = "Treatment effect"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)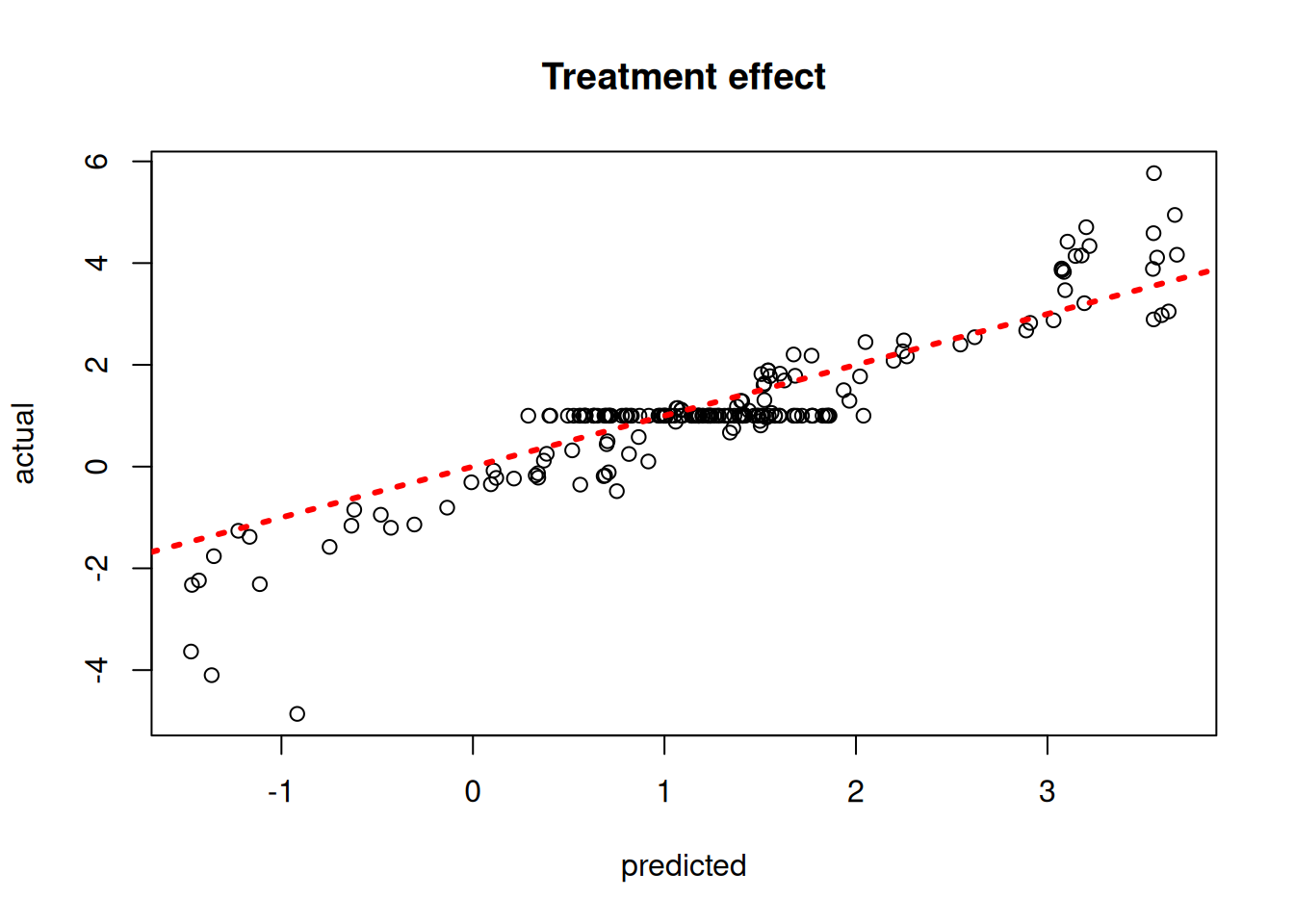
tau_hat_test = bcf_model.predict(X=X_test, Z=Z_test, propensity=pi_test, terms="cate")
tau_pred = tau_hat_test.mean(axis=1)
lo, hi = min(tau_pred.min(), tau_test.min()), max(tau_pred.max(), tau_test.max())
plt.scatter(tau_pred, tau_test, alpha=0.5)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Treatment effect")
plt.show()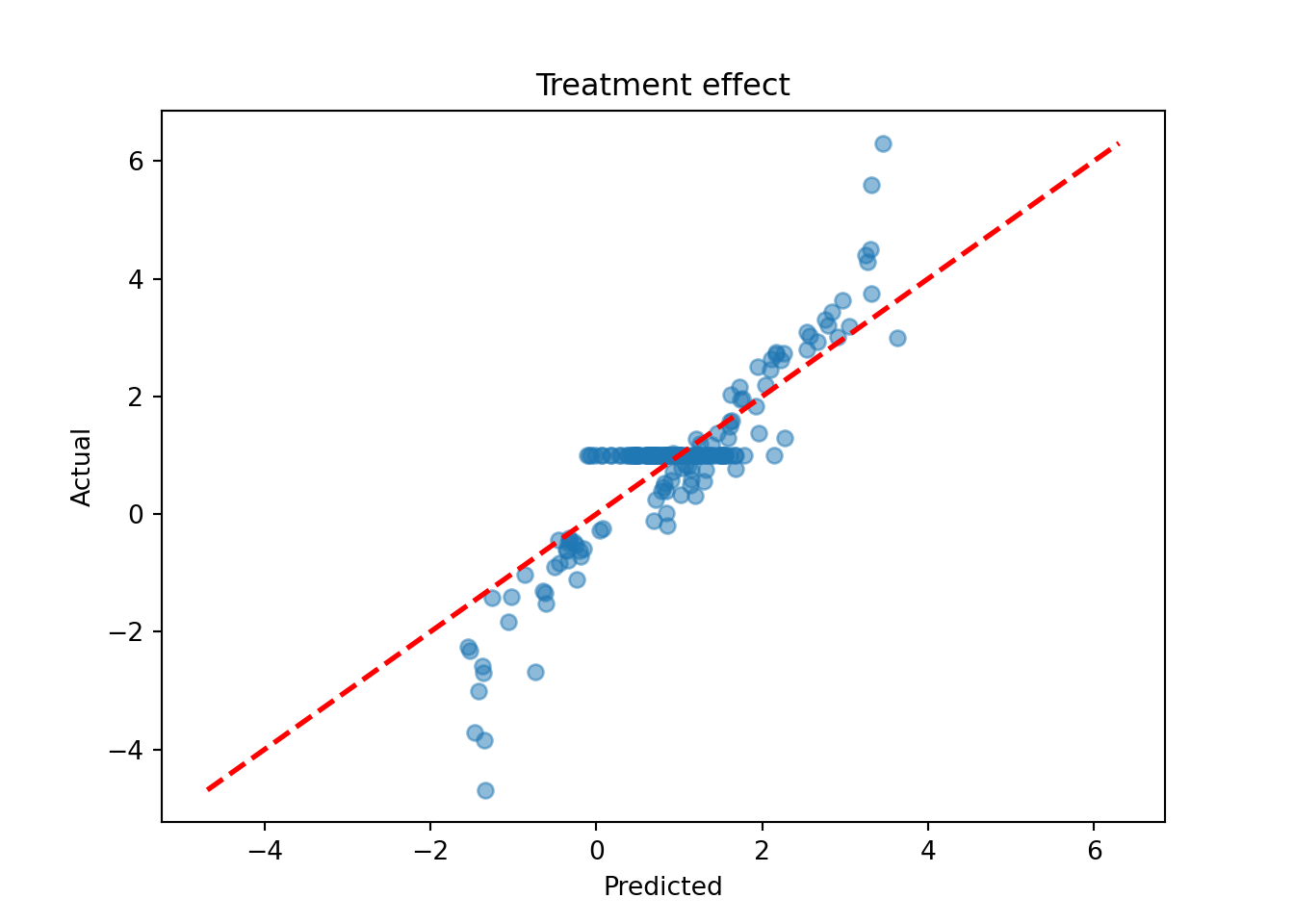
Plot the \(\sigma^2\) traceplot
sigma_observed <- var(y - E_XZ)
sigma2_global_samples <- extractParameter(bcf_model, "sigma2_global")
plot_bounds <- c(
min(c(sigma2_global_samples, sigma_observed)),
max(c(sigma2_global_samples, sigma_observed))
)
plot(
sigma2_global_samples,
ylim = plot_bounds,
ylab = "sigma^2",
xlab = "Sample",
main = "Global variance parameter"
)
abline(h = sigma_observed, lty = 3, lwd = 3, col = "blue")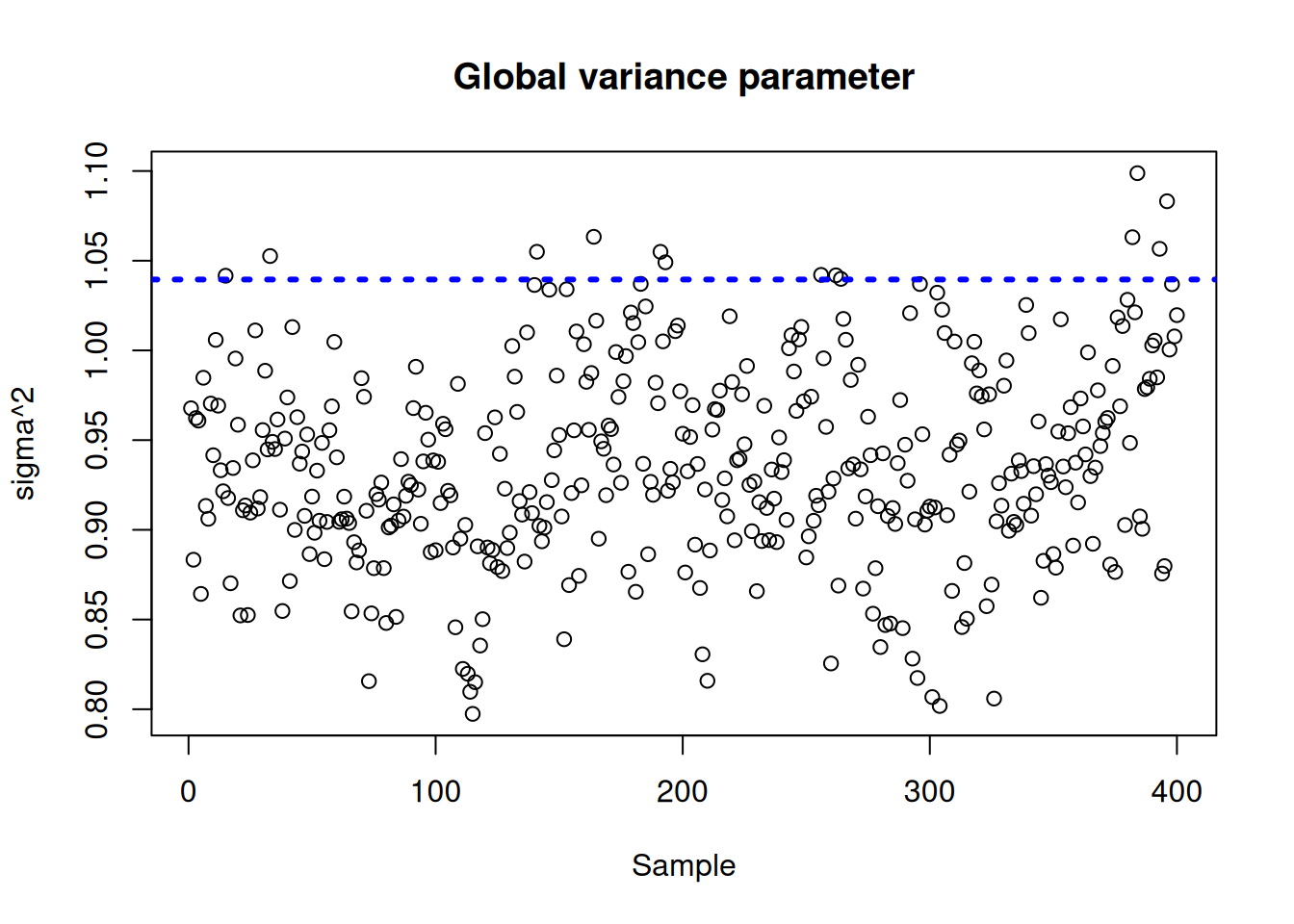
sigma_observed = np.var(y - E_XZ)
global_var_samples = bcf_model.extract_parameter("sigma2_global")
plt.plot(global_var_samples)
plt.axhline(sigma_observed, color="blue", linestyle="dashed", linewidth=2)
plt.xlabel("Sample")
plt.ylabel(r"$\sigma^2$")
plt.title("Global variance parameter")
plt.show()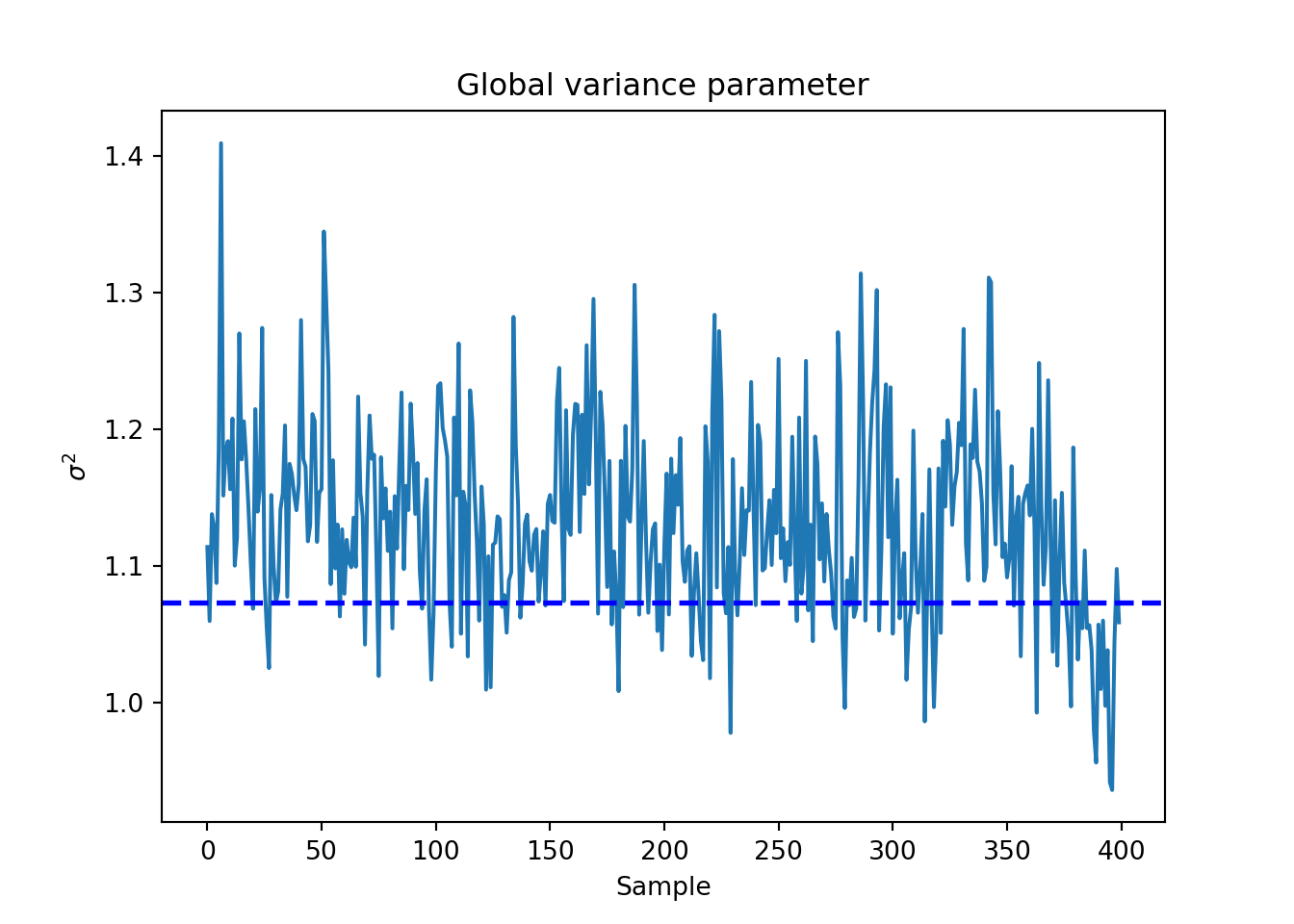
Examine test set interval coverage of \(\tau(X)\).
test_lb <- apply(tau_hat_test, 1, quantile, 0.025)
test_ub <- apply(tau_hat_test, 1, quantile, 0.975)
cover <- ((test_lb <= tau_x[test_inds]) &
(test_ub >= tau_x[test_inds]))
cat("CATE function interval coverage: ", mean(cover) * 100, "%\n")CATE function interval coverage: 84.5 %test_lb = np.quantile(tau_hat_test, 0.025, axis=1)
test_ub = np.quantile(tau_hat_test, 0.975, axis=1)
cover = (test_lb <= tau_test) & (test_ub >= tau_test)
print(f"CATE function interval coverage: {cover.mean() * 100:.2f}%")CATE function interval coverage: 87.50%Demo 2: Nonlinear Outcome Model, Heterogeneous Treatment Effect
We consider the following data generating process from Hahn et al. (2020):
\[\begin{equation*} \begin{aligned} y &= \mu(X) + \tau(X) Z + \epsilon\\ \epsilon &\sim N\left(0,\sigma^2\right)\\ \mu(X) &= 1 + g(X) + 6 \lvert X_3 - 1 \rvert\\ \tau(X) &= 1 + 2 X_2 X_4\\ g(X) &= \mathbb{I}(X_5=1) \times 2 - \mathbb{I}(X_5=2) \times 1 - \mathbb{I}(X_5=3) \times 4\\ s_{\mu} &= \sqrt{\mathbb{V}(\mu(X))}\\ \pi(X) &= 0.8 \phi\left(\frac{3\mu(X)}{s_{\mu}}\right) - \frac{X_1}{2} + \frac{2U+1}{20}\\ X_1,X_2,X_3 &\sim N\left(0,1\right)\\ X_4 &\sim \text{Bernoulli}(1/2)\\ X_5 &\sim \text{Categorical}(1/3,1/3,1/3)\\ U &\sim \text{Uniform}\left(0,1\right)\\ Z &\sim \text{Bernoulli}\left(\pi(X)\right) \end{aligned} \end{equation*}\]
Simulation
Generate data from the DGP above
n <- 1000
snr <- 3
x1 <- rnorm(n)
x2 <- rnorm(n)
x3 <- rnorm(n)
x4 <- as.numeric(rbinom(n, 1, 0.5))
x5 <- as.numeric(sample(1:3, n, replace = TRUE))
X <- cbind(x1, x2, x3, x4, x5)
p <- ncol(X)
mu_x <- mu2(X)
tau_x <- tau2(X)
pi_x <- 0.8 * pnorm((3 * mu_x / sd(mu_x)) - 0.5 * X[, 1]) + 0.05 + runif(n) / 10
Z <- rbinom(n, 1, pi_x)
E_XZ <- mu_x + Z * tau_x
y <- E_XZ + rnorm(n, 0, 1) * (sd(E_XZ) / snr)
X <- as.data.frame(X)
X$x4 <- factor(X$x4, ordered = TRUE)
X$x5 <- factor(X$x5, ordered = TRUE)n = 1000
snr = 3
x1 = rng.normal(size=n)
x2 = rng.normal(size=n)
x3 = rng.normal(size=n)
x4 = rng.binomial(1, 0.5, n).astype(float)
x5 = rng.choice([1, 2, 3], size=n).astype(float)
X = np.column_stack([x1, x2, x3, x4, x5])
mu_x = mu2(X) # mu2 for Demo 2
tau_x = tau2(X)
pi_x = (
0.8 * norm.cdf((3 * mu_x / np.std(mu_x)) - 0.5 * X[:, 0])
+ 0.05
+ rng.uniform(size=n) / 10
)
Z = rng.binomial(1, pi_x, n).astype(float)
E_XZ = mu_x + Z * tau_x
y = E_XZ + rng.normal(size=n) * (np.std(E_XZ) / snr)
X_df = pd.DataFrame({"x1": x1, "x2": x2, "x3": x3, "x4": x4, "x5": x5})
X_df["x4"] = pd.Categorical(X_df["x4"].astype(int), categories=[0, 1], ordered=True)
X_df["x5"] = pd.Categorical(X_df["x5"].astype(int), categories=[1, 2, 3], ordered=True)Split into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct * n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- X[test_inds, ]
X_train <- X[train_inds, ]
pi_test <- pi_x[test_inds]
pi_train <- pi_x[train_inds]
Z_test <- Z[test_inds]
Z_train <- Z[train_inds]
y_test <- y[test_inds]
y_train <- y[train_inds]
mu_test <- mu_x[test_inds]
mu_train <- mu_x[train_inds]
tau_test <- tau_x[test_inds]
tau_train <- tau_x[train_inds]test_set_pct = 0.2
n_test = round(test_set_pct * n)
n_train = n - n_test
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test = X_df.iloc[test_inds]
X_train = X_df.iloc[train_inds]
pi_test, pi_train = pi_x[test_inds], pi_x[train_inds]
Z_test, Z_train = Z[test_inds], Z[train_inds]
y_test, y_train = y[test_inds], y[train_inds]
mu_test, mu_train = mu_x[test_inds], mu_x[train_inds]
tau_test, tau_train = tau_x[test_inds], tau_x[train_inds]Sampling and Analysis
We simulate from a BCF model using default settings.
general_params <- list(
num_threads = 1,
num_chains = 4,
random_seed = random_seed
)
bcf_model <- bcf(
X_train = X_train,
Z_train = Z_train,
y_train = y_train,
propensity_train = pi_train,
X_test = X_test,
Z_test = Z_test,
propensity_test = pi_test,
num_gfr = 10,
num_burnin = 1000,
num_mcmc = 100,
general_params = general_params
)general_params = {"num_threads": 1, "num_chains": 4, "random_seed": random_seed}
bcf_model = BCFModel()
bcf_model.sample(
X_train=X_train,
Z_train=Z_train,
y_train=y_train,
propensity_train=pi_train,
X_test=X_test,
Z_test=Z_test,
propensity_test=pi_test,
num_gfr=10,
num_burnin=1000,
num_mcmc=100,
general_params=general_params,
)Plot the true versus estimated prognostic function
mu_hat_test <- predict(
bcf_model,
X = X_test,
Z = Z_test,
propensity = pi_test,
terms = "prognostic_function"
)
plot(
rowMeans(mu_hat_test),
mu_test,
xlab = "predicted",
ylab = "actual",
main = "Prognostic function"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)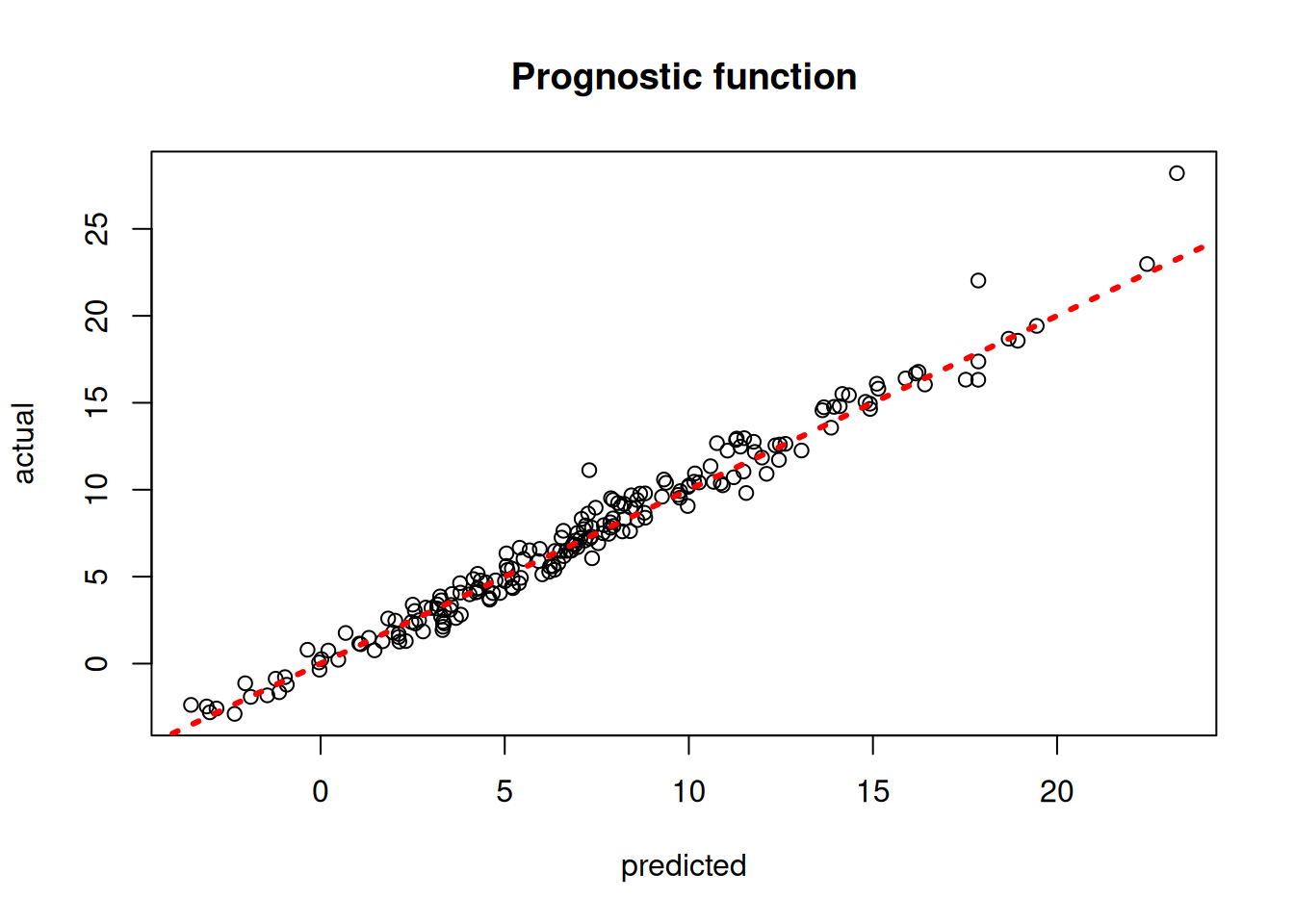
mu_hat_test = bcf_model.predict(
X=X_test, Z=Z_test, propensity=pi_test, terms="prognostic_function"
)
mu_pred = mu_hat_test.mean(axis=1)
lo, hi = min(mu_pred.min(), mu_test.min()), max(mu_pred.max(), mu_test.max())
plt.scatter(mu_pred, mu_test, alpha=0.5)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Prognostic function")
plt.show()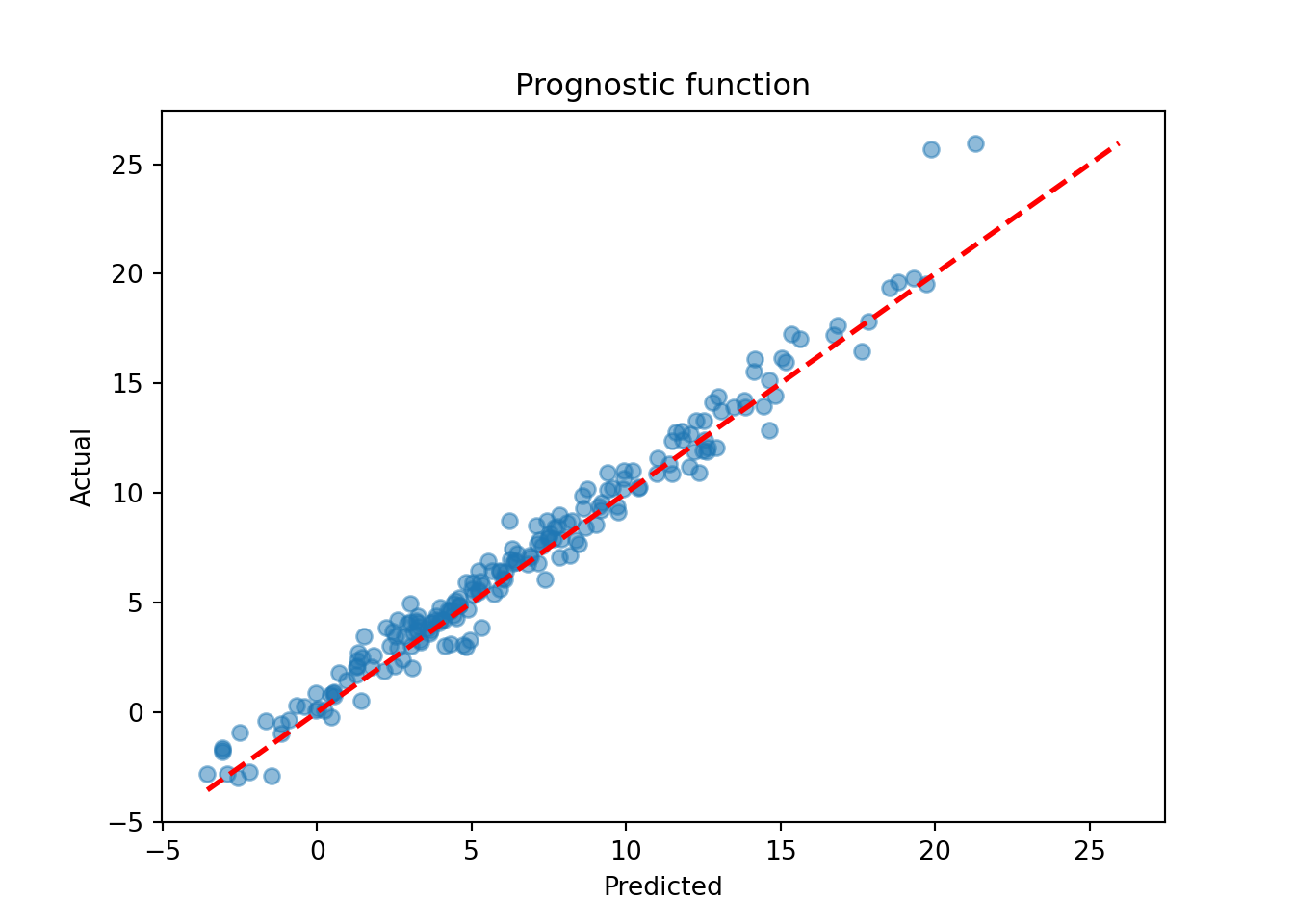
Plot the true versus estimated CATE function
tau_hat_test <- predict(
bcf_model,
X = X_test,
Z = Z_test,
propensity = pi_test,
terms = "cate"
)
plot(
rowMeans(tau_hat_test),
tau_test,
xlab = "predicted",
ylab = "actual",
main = "Treatment effect"
)
abline(0, 1, col = "red", lty = 3, lwd = 3)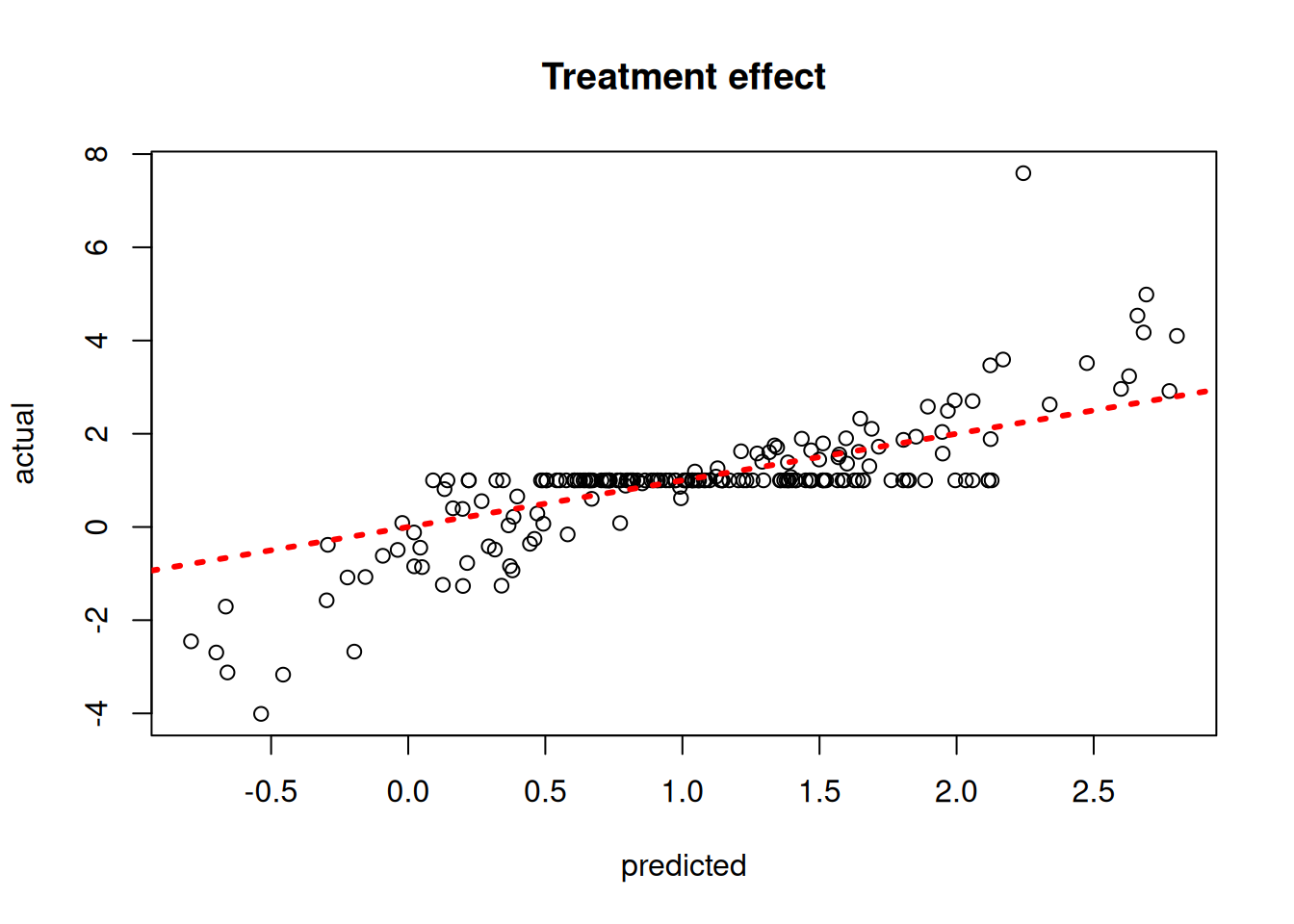
tau_hat_test = bcf_model.predict(X=X_test, Z=Z_test, propensity=pi_test, terms="cate")
tau_pred = tau_hat_test.mean(axis=1)
lo, hi = min(tau_pred.min(), tau_test.min()), max(tau_pred.max(), tau_test.max())
plt.scatter(tau_pred, tau_test, alpha=0.5)
plt.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("Treatment effect")
plt.show()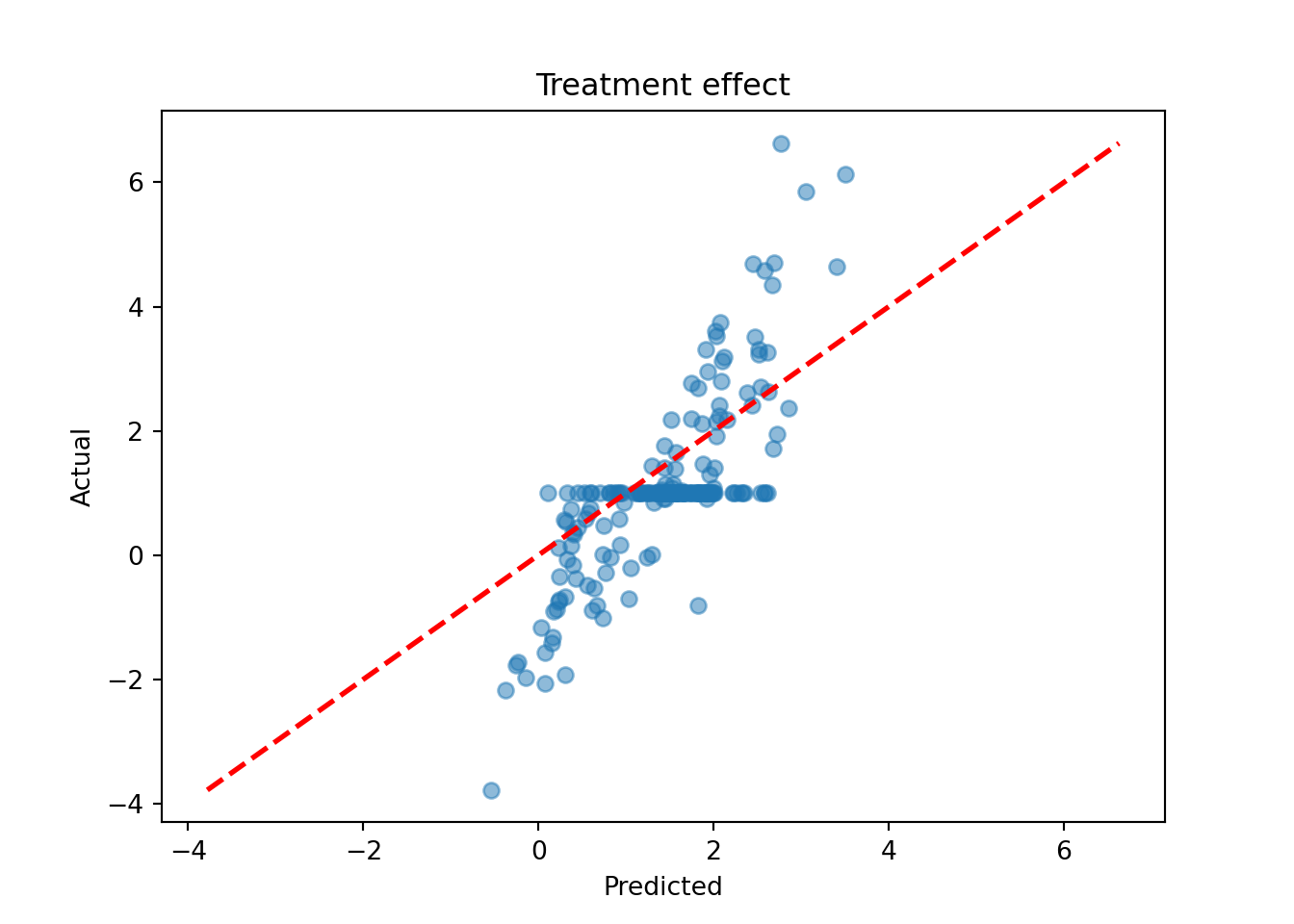
Plot the \(\sigma^2\) traceplot
sigma_observed <- var(y - E_XZ)
sigma2_global_samples <- extractParameter(bcf_model, "sigma2_global")
plot_bounds <- c(
min(c(sigma2_global_samples, sigma_observed)),
max(c(sigma2_global_samples, sigma_observed))
)
plot(
sigma2_global_samples,
ylim = plot_bounds,
ylab = "sigma^2",
xlab = "Sample",
main = "Global variance parameter"
)
abline(h = sigma_observed, lty = 3, lwd = 3, col = "blue")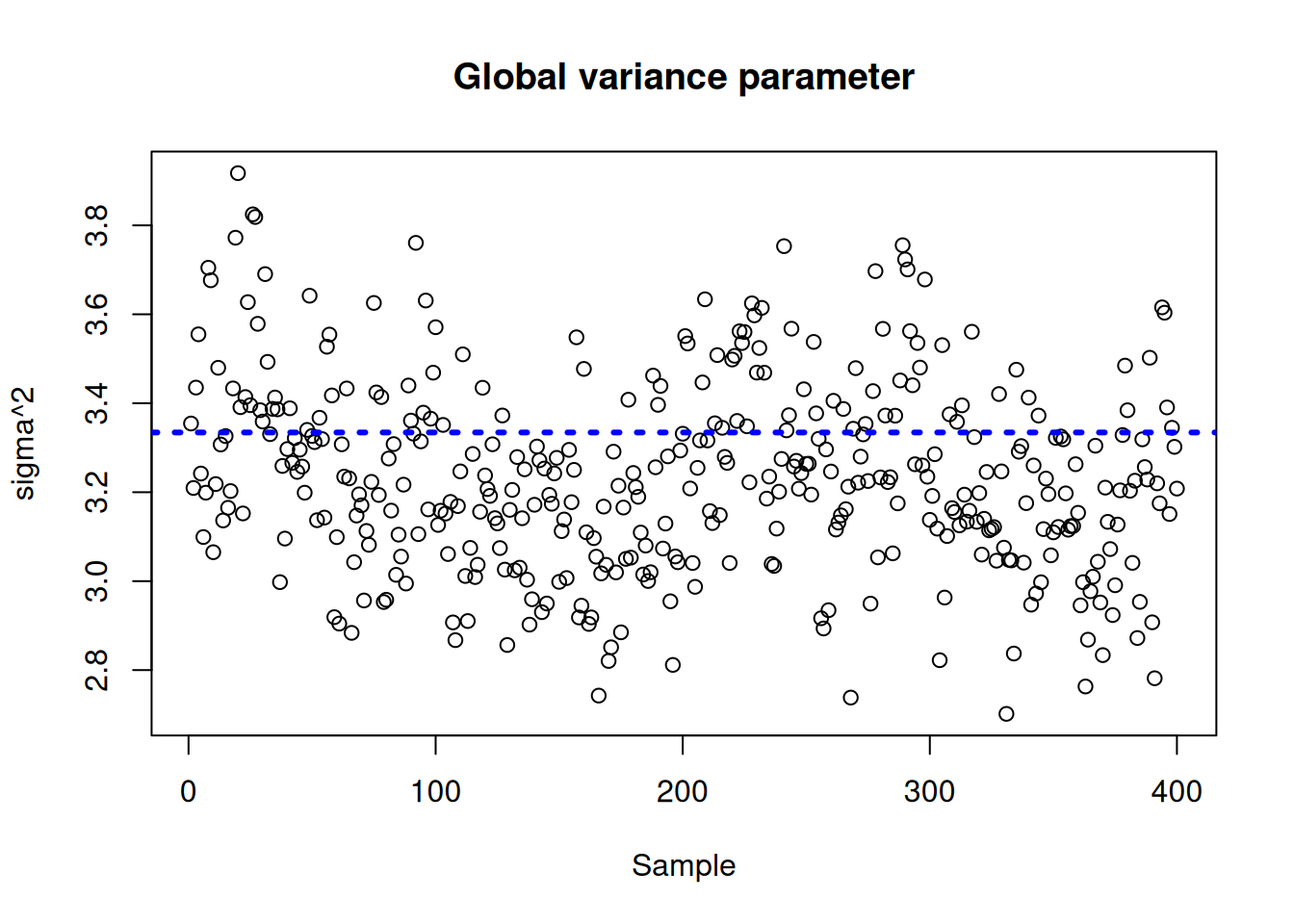
sigma_observed = np.var(y - E_XZ)
global_var_samples = bcf_model.extract_parameter("sigma2_global")
plt.plot(global_var_samples)
plt.axhline(sigma_observed, color="blue", linestyle="dashed", linewidth=2)
plt.xlabel("Sample")
plt.ylabel(r"$\sigma^2$")
plt.title("Global variance parameter")
plt.show()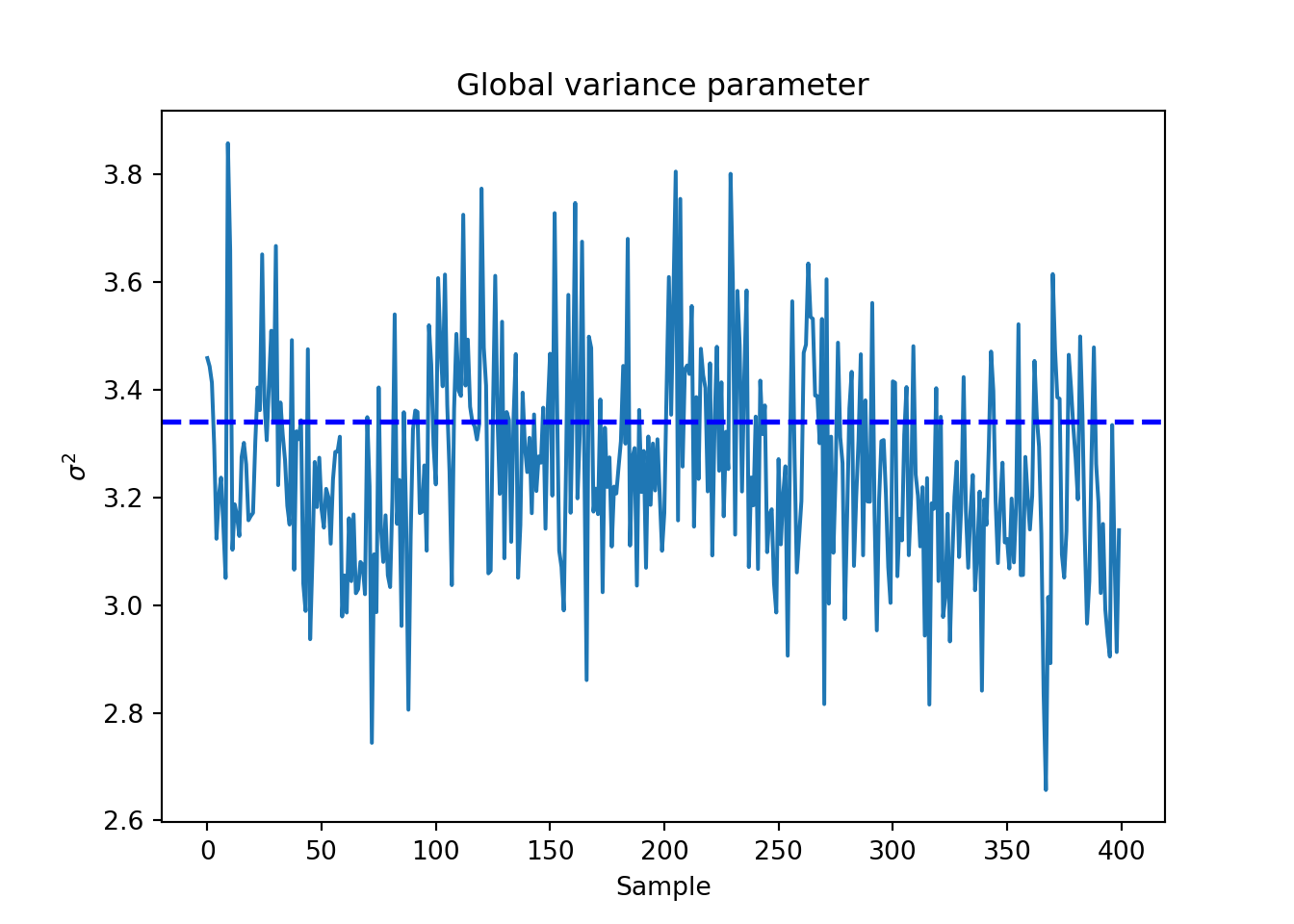
Examine test set interval coverage of \(\tau(X)\).
test_lb <- apply(tau_hat_test, 1, quantile, 0.025)
test_ub <- apply(tau_hat_test, 1, quantile, 0.975)
cover <- ((test_lb <= tau_x[test_inds]) &
(test_ub >= tau_x[test_inds]))
cat("CATE function interval coverage: ", mean(cover) * 100, "%\n")CATE function interval coverage: 92 %test_lb = np.quantile(tau_hat_test, 0.025, axis=1)
test_ub = np.quantile(tau_hat_test, 0.975, axis=1)
cover = (test_lb <= tau_test) & (test_ub >= tau_test)
print(f"CATE function interval coverage: {cover.mean() * 100:.2f}%")CATE function interval coverage: 84.00%