library(stochtree)Posterior Summary and Visualization Utilities
This vignette demonstrates the summary and plotting utilities available for stochtree models.
Setup
Load necessary packages
import numpy as np
import matplotlib.pyplot as plt
from stochtree import BARTModel, BCFModel, plot_parameter_traceSet a seed for reproducibility
random_seed = 1234
set.seed(random_seed)random_seed = 1234
rng = np.random.default_rng(random_seed)Supervised Learning
We begin with the supervised learning use case served by the bart() function.
Below we simulate a simple regression dataset.
n <- 1000
p_x <- 10
p_w <- 1
X <- matrix(runif(n * p_x), ncol = p_x)
W <- matrix(runif(n * p_w), ncol = p_w)
f_XW <- (((0 <= X[, 10]) & (0.25 > X[, 10])) *
(-7.5 * W[, 1]) +
((0.25 <= X[, 10]) & (0.5 > X[, 10])) * (-2.5 * W[, 1]) +
((0.5 <= X[, 10]) & (0.75 > X[, 10])) * (2.5 * W[, 1]) +
((0.75 <= X[, 10]) & (1 > X[, 10])) * (7.5 * W[, 1]))
noise_sd <- 1
y <- f_XW + rnorm(n, 0, 1) * noise_sdn = 1000
p_x = 10
p_w = 1
X = rng.uniform(size=(n, p_x))
W = rng.uniform(size=(n, p_w))
# R uses X[,10] (1-indexed) = Python X[:,9]
f_XW = (
((X[:, 9] >= 0) & (X[:, 9] < 0.25)) * (-7.5 * W[:, 0]) +
((X[:, 9] >= 0.25) & (X[:, 9] < 0.5)) * (-2.5 * W[:, 0]) +
((X[:, 9] >= 0.5) & (X[:, 9] < 0.75)) * ( 2.5 * W[:, 0]) +
((X[:, 9] >= 0.75) & (X[:, 9] < 1.0)) * ( 7.5 * W[:, 0])
)
noise_sd = 1.0
y = f_XW + rng.standard_normal(n) * noise_sdNow we fit a simple BART model to the data.
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 1000
general_params <- list(
num_threads = 1,
num_chains = 3
)
bart_model <- stochtree::bart(
X_train = X,
y_train = y,
leaf_basis_train = W,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params
)bart_model = BARTModel()
bart_model.sample(
X_train=X,
y_train=y,
leaf_basis_train=W,
num_gfr=10,
num_burnin=0,
num_mcmc=1000,
general_params={
"num_threads": 1,
"num_chains": 3
},
)We obtain a high level summary of the BART model by running print().
print(bart_model)stochtree::bart() run with mean forest, global error variance model, and mean forest leaf scale model
Continuous outcome was modeled as Gaussian with a leaf regression prior with 1 bases for the mean forest
Outcome was standardized
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning) print(bart_model)BARTModel run with mean forest, global error variance model, and mean forest leaf scale model
Outcome was modeled as gaussian with a leaf regression prior with 1 bases for the mean forest
Outcome was standardized
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)For a more detailed summary (including the information above), we use the summary() function.
summary(bart_model)stochtree::bart() run with mean forest, global error variance model, and mean forest leaf scale model
Continuous outcome was modeled as Gaussian with a leaf regression prior with 1 bases for the mean forest
Outcome was standardized
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)
Summary of sigma^2 posterior:
3000 samples, mean = 0.888, standard deviation = 0.049, quantiles:
2.5% 10% 25% 50% 75% 90% 97.5%
0.7976738 0.8276973 0.8531417 0.8850262 0.9191620 0.9512392 0.9905685
Summary of leaf scale posterior:
3000 samples, mean = 0.007, standard deviation = 0.001, quantiles:
2.5% 10% 25% 50% 75% 90%
0.005152223 0.005820257 0.006539487 0.007328878 0.008172267 0.009171917
97.5%
0.010802867
Summary of in-sample posterior mean predictions:
1000 observations, mean = -0.064, standard deviation = 3.286, quantiles:
2.5% 10% 25% 50% 75% 90% 97.5%
-6.8485115 -4.9269938 -1.8268155 -0.1202225 2.0285138 4.2590778 6.5453792 print(bart_model.summary())BART Model Summary:
-------------------
BARTModel run with mean forest, global error variance model, and mean forest leaf scale model
Outcome was modeled as gaussian with a leaf regression prior with 1 bases for the mean forest
Outcome was standardized
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)
Summary of sigma^2 posterior: 3000 samples, mean = 0.934, standard deviation = 0.049, quantiles:
2.5%: 0.845
10.0%: 0.872
25.0%: 0.901
50.0%: 0.933
75.0%: 0.966
90.0%: 0.998
97.5%: 1.037
Summary of leaf scale posterior: 3000 samples, mean = 0.007, standard deviation = 0.001, quantiles:
2.5%: 0.005
10.0%: 0.005
25.0%: 0.006
50.0%: 0.007
75.0%: 0.007
90.0%: 0.008
97.5%: 0.009
Summary of in-sample posterior mean predictions:
1000 observations, mean = 0.106, standard deviation = 3.286, quantiles:
2.5%: -6.737
10.0%: -4.313
25.0%: -1.950
50.0%: 0.169
75.0%: 2.093
90.0%: 4.420
97.5%: 6.575
NoneWe can use the plot() function to produce a traceplot of model terms like the global error scale \(\sigma^2\) or (if \(\sigma^2\) is not sampled) the first observation of cached train set predictions.
plot(bart_model)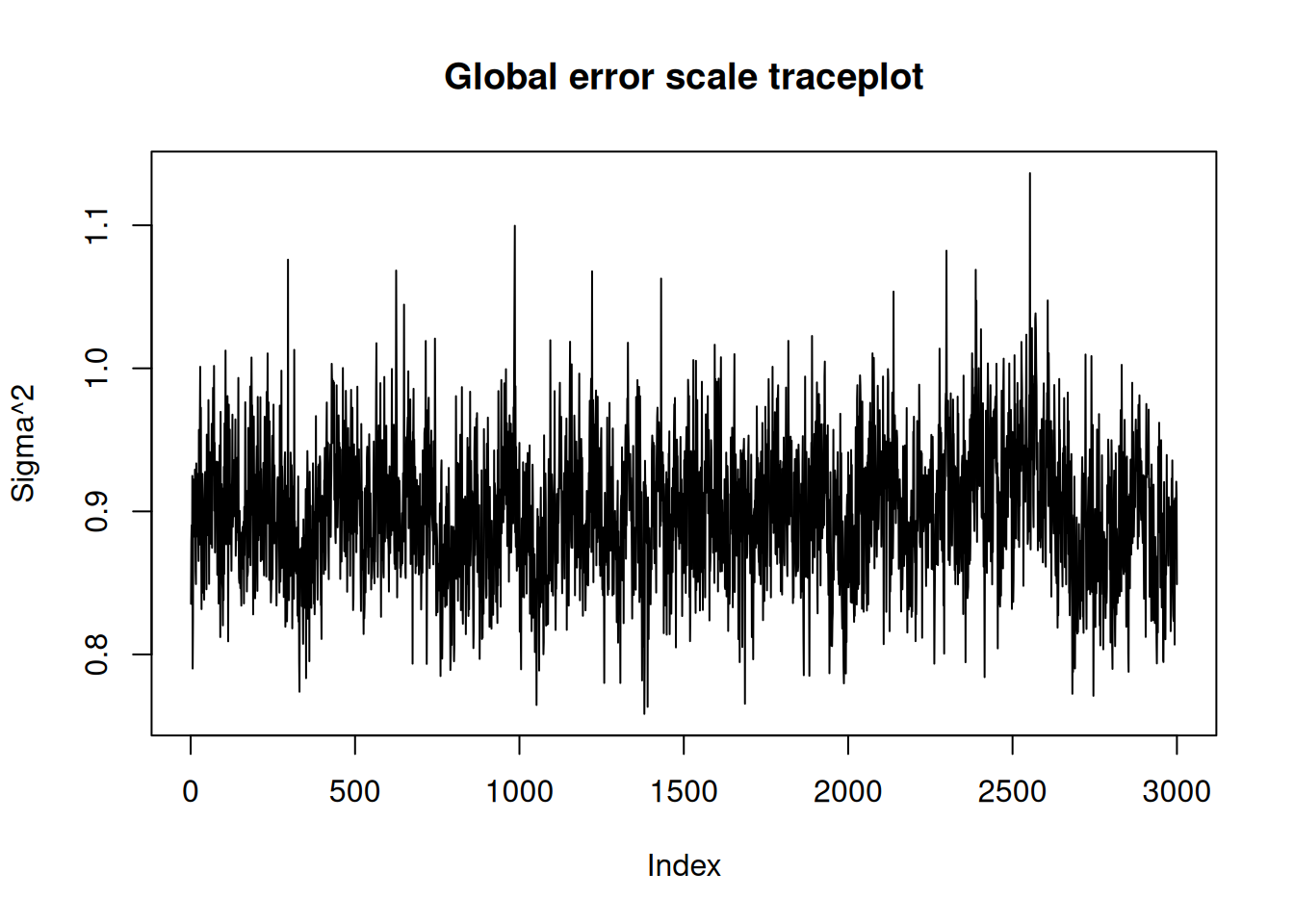
ax = plot_parameter_trace(bart_model, term="global_error_scale")
plt.show()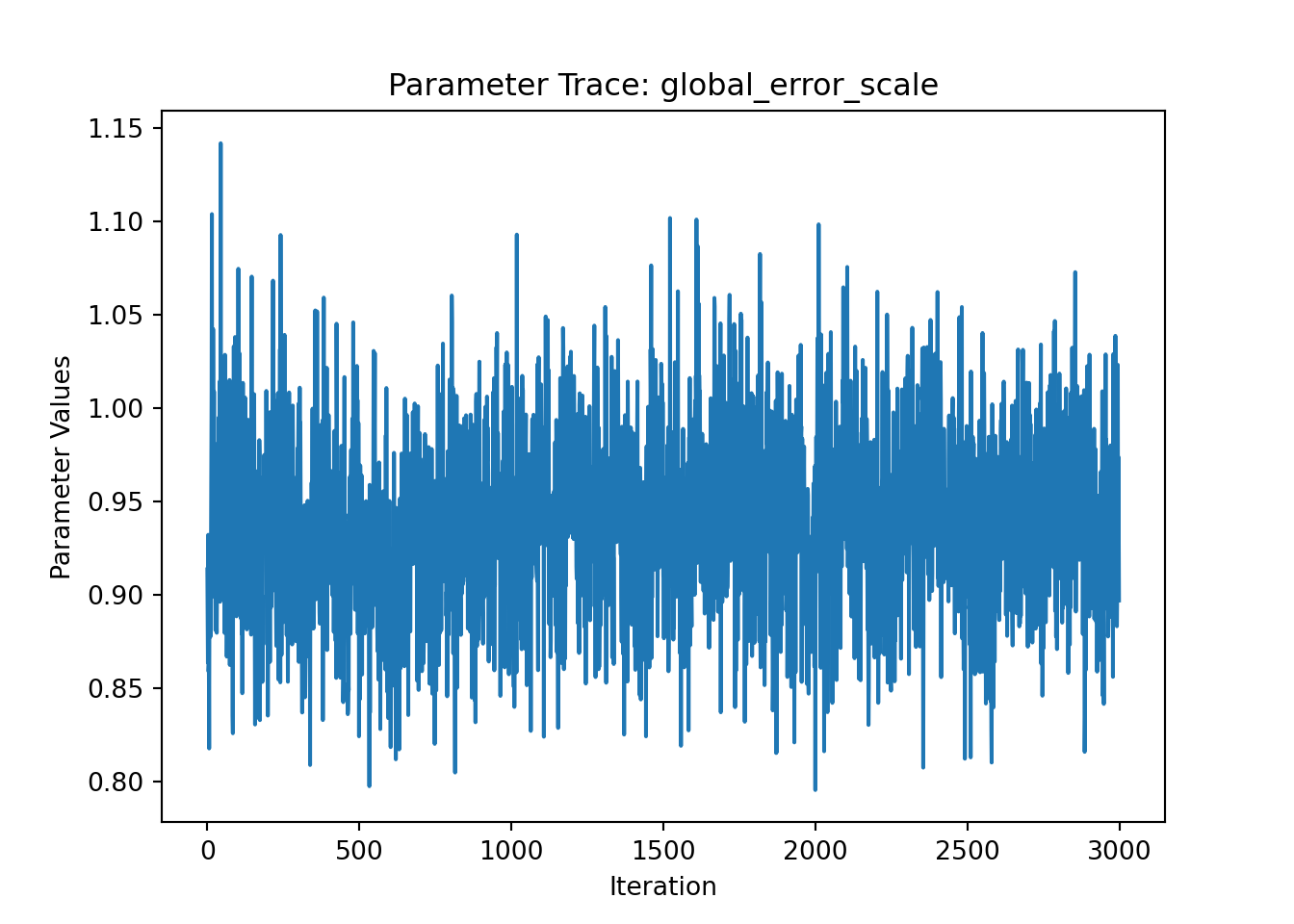
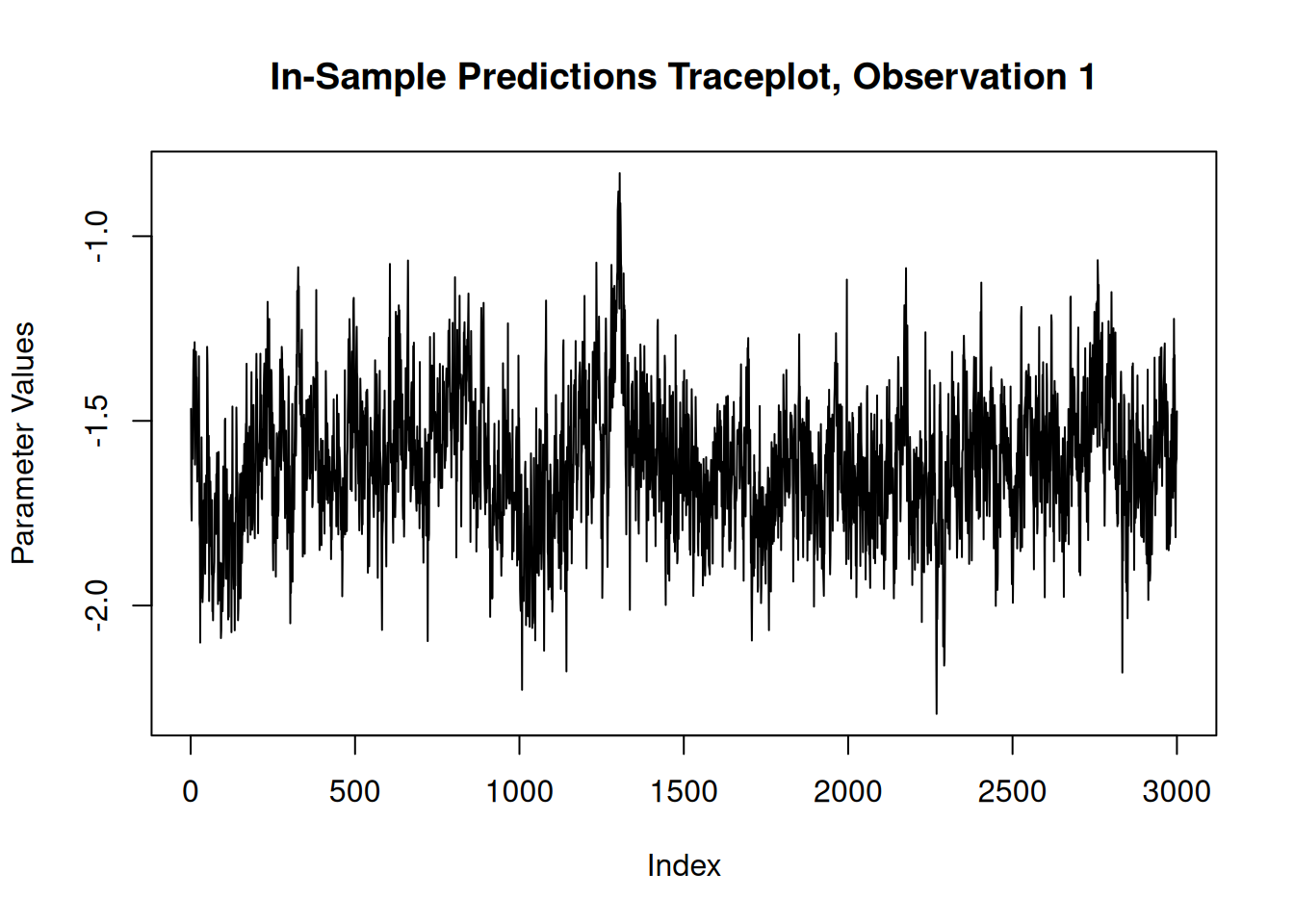
For finer-grained control over which parameters to plot, we can also use the extractParameter() function to pull the posterior distribution of any valid model term (e.g., global error scale \(\sigma^2\), leaf scale \(\sigma^2_{\ell}\), in-sample mean function predictions y_hat_train) and then plot any subset or transformation of these values.
y_hat_train_samples <- extractParameter(bart_model, "y_hat_train")
obs_index <- 1
plot(
y_hat_train_samples[obs_index, ],
type = "l",
main = paste0("In-Sample Predictions Traceplot, Observation ", obs_index),
xlab = "Index",
ylab = "Parameter Values"
)
y_hat_train_samples = bart_model.extract_parameter("y_hat_train")
obs_index = 0
fig, ax = plt.subplots()
ax.plot(y_hat_train_samples[obs_index, :])
ax.set_title(f"In-Sample Predictions Traceplot, Observation {obs_index}")
ax.set_xlabel("Index")
ax.set_ylabel("Parameter Values")
plt.show()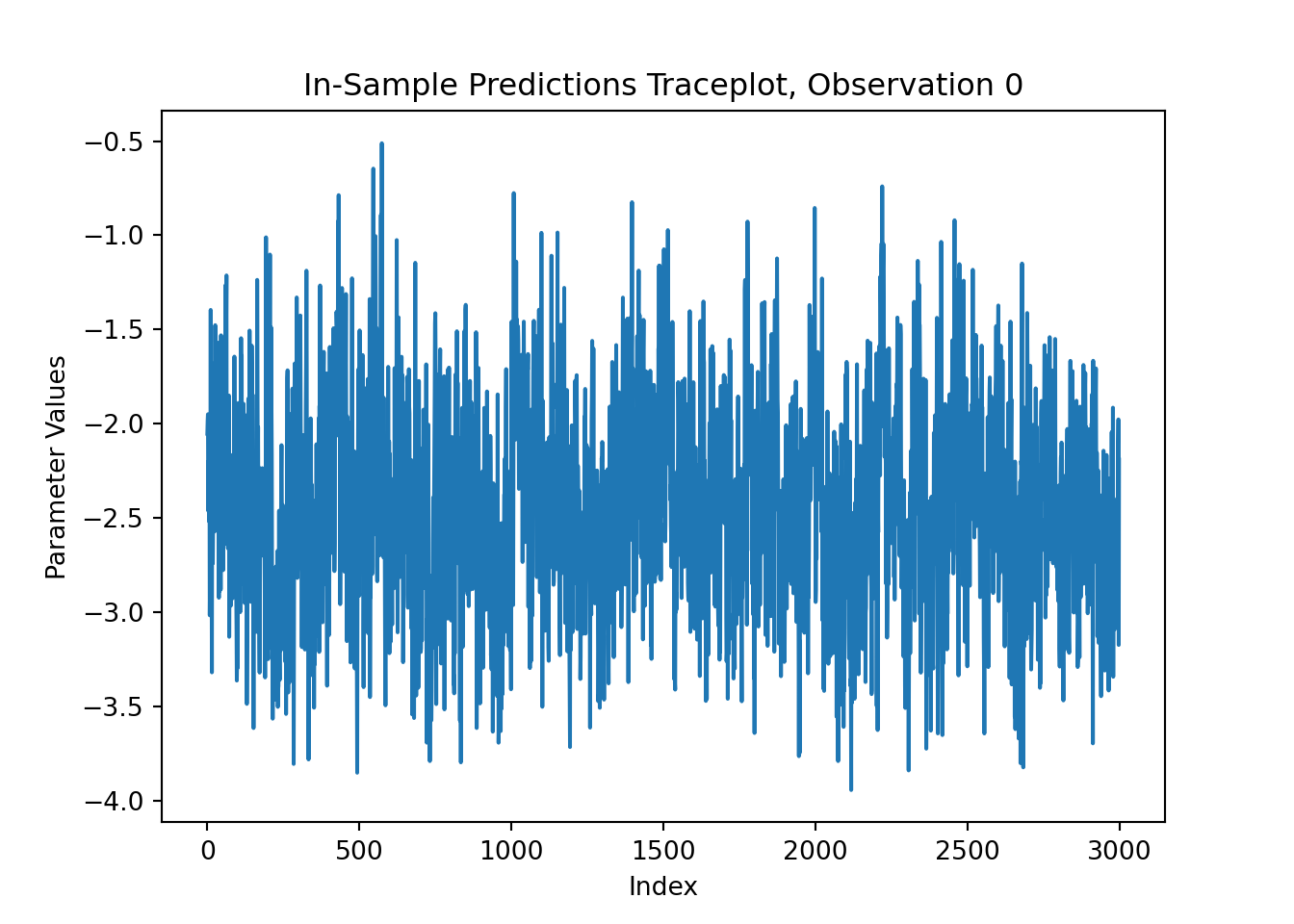
Causal Inference
We now run the same demo for the causal inference use case served by the bcf() function in R and the BCFModel Python class.
Below we simulate a simple dataset for a causal inference problem with binary treatment and continuous outcome.
# Generate covariates and treatment
n <- 1000
p_X = 5
X = matrix(runif(n * p_X), ncol = p_X)
pi_X = 0.25 + 0.5 * X[, 1]
Z = rbinom(n, 1, pi_X)
# Define the outcome mean functions (prognostic and treatment effects)
mu_X = pi_X * 5 + 2 * X[, 3]
tau_X = X[, 2] * 2 - 1
# Generate outcome
epsilon = rnorm(n, 0, 1)
y = mu_X + tau_X * Z + epsilon# Generate covariates and treatment
n = 1000
p_X = 5
X = rng.uniform(size=(n, p_X))
pi_X = 0.25 + 0.5 * X[:, 0]
Z = rng.binomial(1, pi_X, n).astype(float)
# Define the outcome mean functions (prognostic and treatment effects)
mu_X = pi_X * 5 + 2 * X[:, 2]
tau_X = X[:, 1] * 2 - 1
# Generate outcome
epsilon = rng.standard_normal(n)
y = mu_X + tau_X * Z + epsilonNow we fit a simple BCF model to the data
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 1000
general_params <- list(
num_threads = 1,
num_chains = 3,
adaptive_coding = TRUE
)
bcf_model <- stochtree::bcf(
X_train = X,
y_train = y,
Z_train = Z,
num_gfr = num_gfr,
num_burnin = num_burnin,
num_mcmc = num_mcmc,
general_params = general_params
)bcf_model = BCFModel()
bcf_model.sample(
X_train=X,
Z_train=Z,
y_train=y,
propensity_train=pi_X,
num_gfr=10,
num_burnin=0,
num_mcmc=1000,
general_params={
"num_threads": 1,
"num_chains": 3,
"adaptive_coding": True
},
)We obtain a high level summary of the BCF model by running print().
print(bcf_model)stochtree::bcf() run with prognostic forest, treatment effect forest, global error variance model, prognostic forest leaf scale model, and treatment effect intercept model
Outcome was modeled as gaussian
Treatment was binary and its effect was estimated with adaptive coding
outcome was standardized
An internal propensity model was fit using stochtree::bart() in lieu of user-provided propensity scores
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning) print(bcf_model)BCFModel run with prognostic forest, treatment effect forest, global error variance model, prognostic forest leaf scale model, and treatment effect intercept model
Outcome was modeled as gaussian
Treatment was binary and its effect was estimated with adaptive coding
Outcome was standardized
User-provided propensity scores were included in the model
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)For a more detailed summary (including the information above), we use the summary() function / method.
summary(bcf_model)stochtree::bcf() run with prognostic forest, treatment effect forest, global error variance model, prognostic forest leaf scale model, and treatment effect intercept model
Outcome was modeled as gaussian
Treatment was binary and its effect was estimated with adaptive coding
outcome was standardized
An internal propensity model was fit using stochtree::bart() in lieu of user-provided propensity scores
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)
Summary of sigma^2 posterior:
3000 samples, mean = 0.949, standard deviation = 0.045, quantiles:
2.5% 10% 25% 50% 75% 90% 97.5%
0.8643862 0.8907757 0.9182082 0.9484272 0.9783383 1.0078629 1.0409780
Summary of prognostic forest leaf scale posterior:
3000 samples, mean = 0.002, standard deviation = 0.000, quantiles:
2.5% 10% 25% 50% 75% 90%
0.0008568294 0.0010371220 0.0012421974 0.0014786088 0.0018064272 0.0021478841
97.5%
0.0025004825
Summary of adaptive coding parameters:
3000 samples, mean (control) = -0.437, mean (treated) = 1.029, standard deviation (control) = 0.348, standard deviation (treated) = 0.315
quantiles (control):
2.5% 10% 25% 50% 75% 90%
-1.17874376 -0.89652527 -0.66059102 -0.41437907 -0.18414640 -0.01349886
97.5%
0.16926139
quantiles (treated):
2.5% 10% 25% 50% 75% 90% 97.5%
0.4624619 0.6462934 0.8138740 1.0075352 1.2248498 1.4620051 1.6881949
Summary of treatment effect intercept (tau_0) posterior:
3000 samples, mean = 0.164, standard deviation = 0.734, quantiles:
2.5% 10% 25% 50% 75% 90% 97.5%
-1.3587447 -0.9591607 -0.2164569 0.1000003 0.7793058 1.1575356 1.3306564
Summary of in-sample posterior mean predictions:
1000 observations, mean = 3.487, standard deviation = 0.946, quantiles:
2.5% 10% 25% 50% 75% 90% 97.5%
1.654992 2.250739 2.792019 3.475277 4.131819 4.727274 5.379340
Summary of in-sample posterior mean CATEs:
1000 observations, mean = 0.063, standard deviation = 0.524, quantiles:
2.5% 10% 25% 50% 75% 90%
-0.73251543 -0.55922780 -0.39695284 -0.03452641 0.57664883 0.79384361
97.5%
0.91214143 print(bcf_model.summary())BCF Model Summary:
------------------
BCFModel run with prognostic forest, treatment effect forest, global error variance model, prognostic forest leaf scale model, and treatment effect intercept model
Outcome was modeled as gaussian
Treatment was binary and its effect was estimated with adaptive coding
Outcome was standardized
User-provided propensity scores were included in the model
The sampler was run for 10 GFR iterations, with 3 chains of 0 burn-in iterations and 1000 MCMC iterations, retaining every iteration (i.e. no thinning)
Summary of sigma^2 posterior: 3000 samples, mean = 0.872, standard deviation = 0.042, quantiles:
2.5%: 0.794
10.0%: 0.819
25.0%: 0.843
50.0%: 0.871
75.0%: 0.900
90.0%: 0.924
97.5%: 0.958
Summary of prognostic forest leaf scale posterior: 3000 samples, mean = 0.002, standard deviation = 0.001, quantiles:
2.5%: 0.001
10.0%: 0.001
25.0%: 0.001
50.0%: 0.002
75.0%: 0.002
90.0%: 0.002
97.5%: 0.003
Summary of adaptive coding parameters:
3000 samples, mean (control) = -0.575, mean (treated) = 1.114, standard deviation (control) = 0.365, standard deviation (treated) = 0.352
quantiles (control):
2.5%: -1.346
10.0%: -1.053
25.0%: -0.801
50.0%: -0.547
75.0%: -0.324
90.0%: -0.147
97.5%: 0.093
quantiles (treated):
2.5%: 0.501
10.0%: 0.672
25.0%: 0.875
50.0%: 1.096
75.0%: 1.330
90.0%: 1.547
97.5%: 1.868
Summary of treatment effect intercept (tau_0) posterior: 3000 samples, mean = 0.025, standard deviation = 0.584, quantiles:
2.5%: -1.121
10.0%: -0.909
25.0%: -0.285
50.0%: 0.087
75.0%: 0.475
90.0%: 0.737
97.5%: 0.955
Summary of in-sample posterior mean predictions:
1000 observations, mean = 3.482, standard deviation = 0.961, quantiles:
2.5%: 1.854
10.0%: 2.210
25.0%: 2.779
50.0%: 3.475
75.0%: 4.157
90.0%: 4.775
97.5%: 5.334
Summary of in-sample posterior mean CATEs:
1000 observations, mean = -0.028, standard deviation = 0.625, quantiles:
2.5%: -1.071
10.0%: -0.912
25.0%: -0.587
50.0%: 0.071
75.0%: 0.540
90.0%: 0.741
97.5%: 0.920
NoneIn R, we have a plot() that produces a traceplot of model terms like the global error scale \(\sigma^2\) or (if \(\sigma^2\) is not sampled) the first observation of cached train set predictions.
In Python, we provide a plot_parameter_trace() function for requesting a traceplot of a specific model parameter.
plot(bcf_model)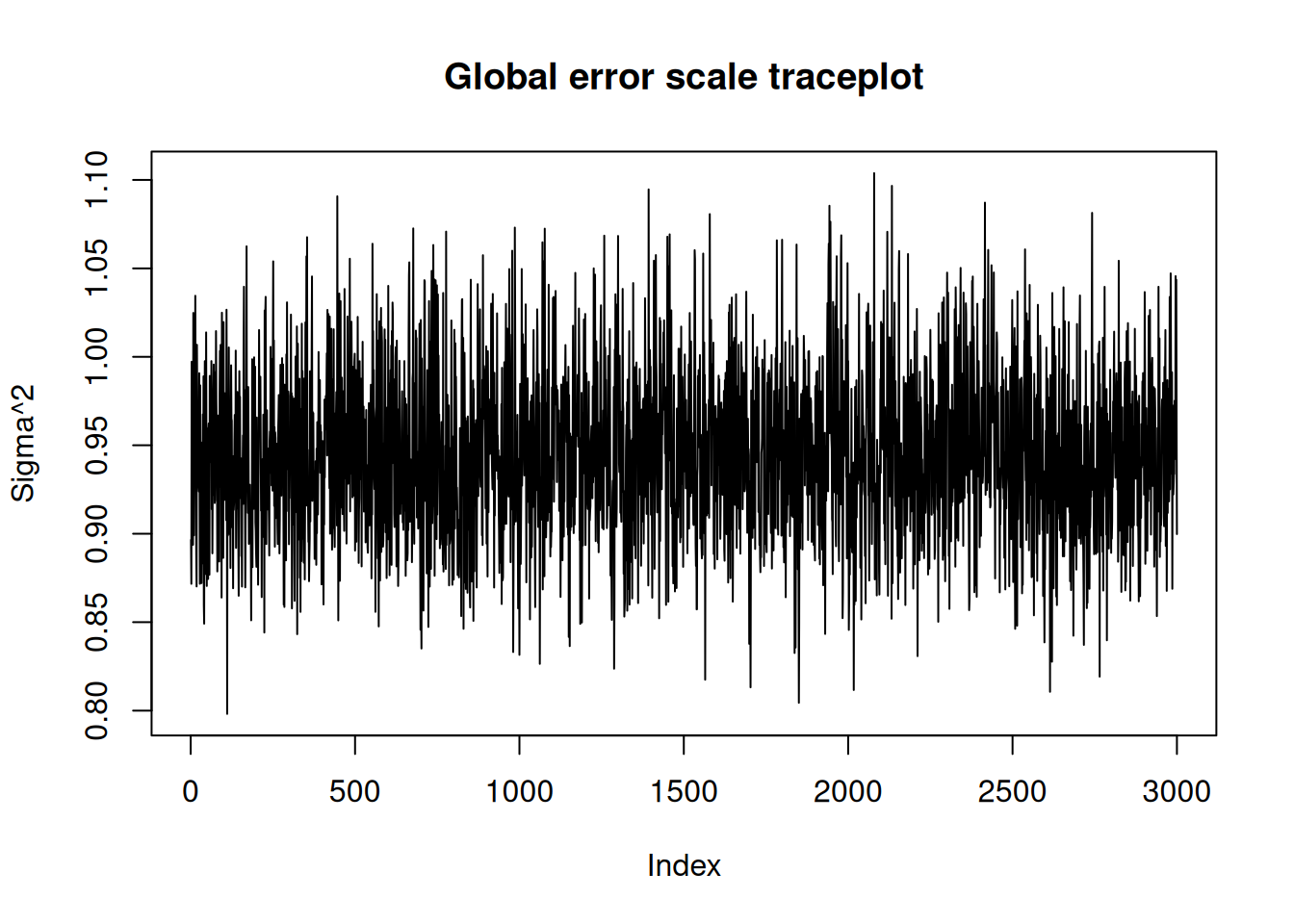
ax = plot_parameter_trace(bcf_model, term="global_error_scale")
plt.show()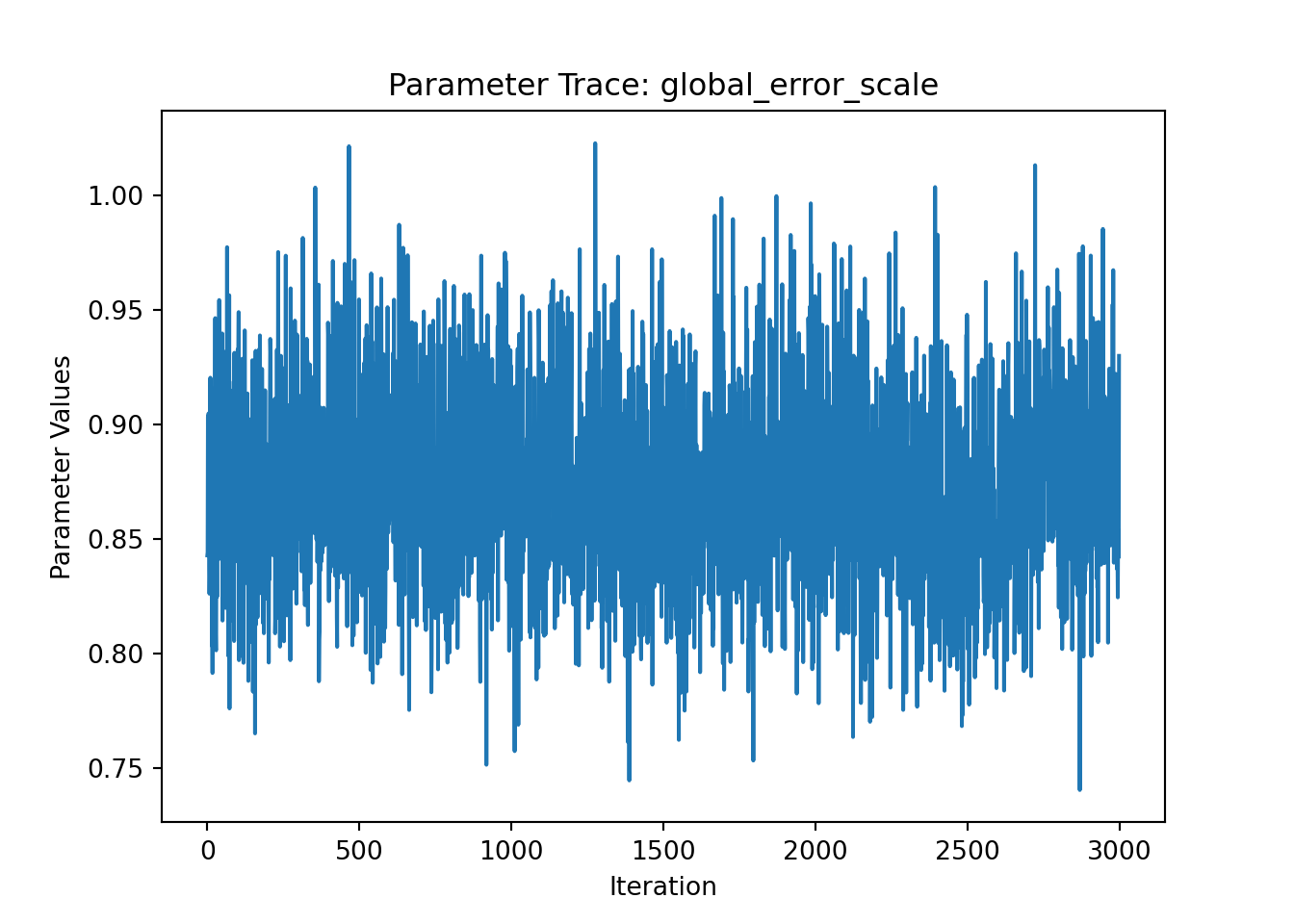
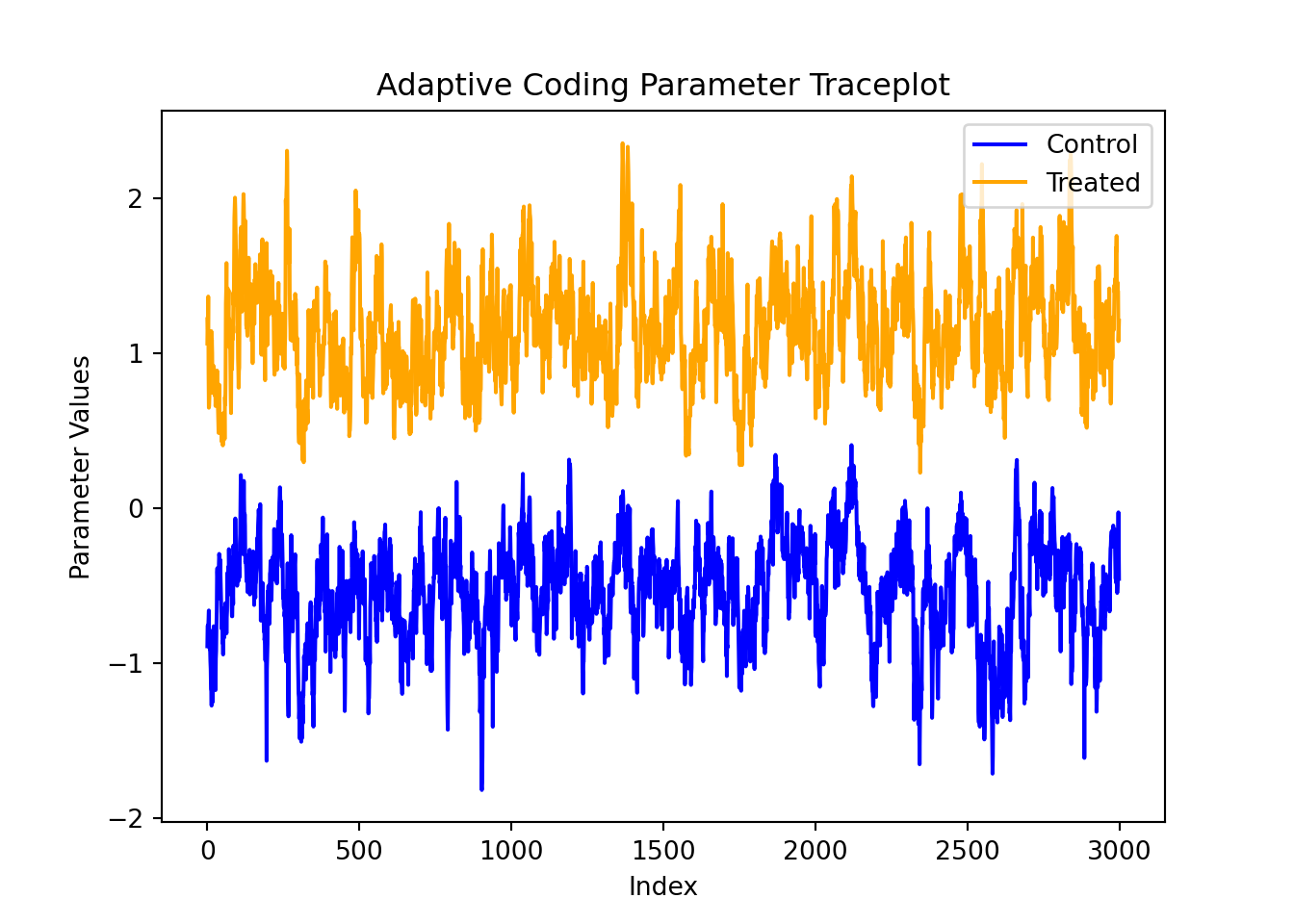
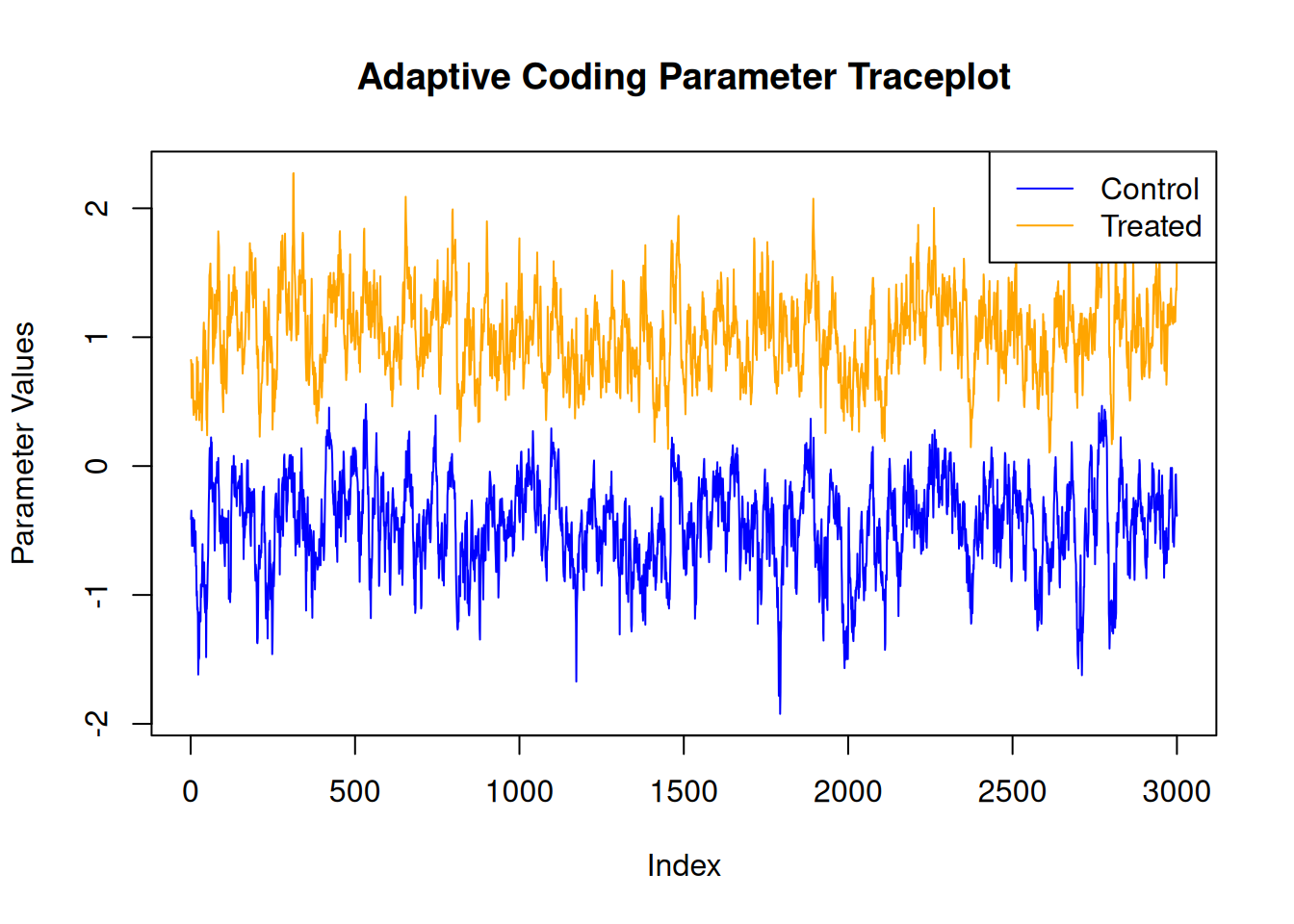
For finer-grained control over which parameters to plot, we can also use the extractParameter() function in R or the extract_parameter() method in Python to query the posterior distribution of any valid model term (e.g., global error scale \(\sigma^2\), prognostic forest leaf scale \(\sigma^2_{\mu}\), CATE forest leaf scale \(\sigma^2_{\tau}\), adaptive coding parameters \(b_0\) and \(b_1\) for binary treatment, in-sample mean function predictions y_hat_train, in-sample CATE function predictions tau_hat_train) and then plot any subset or transformation of these values.
adaptive_coding_samples <- extractParameter(bcf_model, "adaptive_coding")
plot(
adaptive_coding_samples[1, ],
type = "l",
main = "Adaptive Coding Parameter Traceplot",
xlab = "Index",
ylab = "Parameter Values",
ylim = range(adaptive_coding_samples),
col = "blue"
)
lines(adaptive_coding_samples[2, ], col = "orange")
legend(
"topright",
legend = c("Control", "Treated"),
lty = 1,
col = c("blue", "orange")
)
adaptive_coding_samples = bcf_model.extract_parameter("adaptive_coding")
fig, ax = plt.subplots()
ax.plot(adaptive_coding_samples[0, :], color="blue", label="Control")
ax.plot(adaptive_coding_samples[1, :], color="orange", label="Treated")
ax.set_title("Adaptive Coding Parameter Traceplot")
ax.set_xlabel("Index")
ax.set_ylabel("Parameter Values")
ax.legend(loc="upper right")
plt.show()