library(stochtree)Examining Individual Trees in a Fitted Ensemble
While out of sample evaluation and MCMC diagnostics on parametric BART components (i.e. \(\sigma^2\), the global error variance) are helpful, it’s important to be able to inspect the trees in a BART / BCF model. This vignette walks through some of the features stochtree provides to query and understand the forests and trees in a model.
Setup
Load necessary packages
import numpy as np
import matplotlib.pyplot as plt
from stochtree import BARTModelSet a seed for reproducibility
random_seed = 1234
set.seed(random_seed)random_seed = 1234
rng = np.random.default_rng(random_seed)Data Generation
Generate sample data where feature 10 is the only “important” feature
n <- 500
p_x <- 10
X <- matrix(runif(n*p_x), ncol = p_x)
f_XW <- (
((0 <= X[,10]) & (0.25 > X[,10])) * (-7.5) +
((0.25 <= X[,10]) & (0.5 > X[,10])) * (-2.5) +
((0.5 <= X[,10]) & (0.75 > X[,10])) * (2.5) +
((0.75 <= X[,10]) & (1 > X[,10])) * (7.5)
)
noise_sd <- 1
y <- f_XW + rnorm(n, 0, 1)*noise_sdn = 500
p_x = 10
X = rng.uniform(size=(n, p_x))
# Feature 10 (R) = feature index 9 (Python, 0-indexed)
f_XW = (
((X[:, 9] >= 0) & (X[:, 9] < 0.25)) * (-7.5) +
((X[:, 9] >= 0.25) & (X[:, 9] < 0.5)) * (-2.5) +
((X[:, 9] >= 0.5) & (X[:, 9] < 0.75)) * (2.5) +
((X[:, 9] >= 0.75) & (X[:, 9] < 1.0)) * (7.5)
)
noise_sd = 1.0
y = f_XW + rng.standard_normal(n) * noise_sdSplit into train and test sets
test_set_pct <- 0.2
n_test <- round(test_set_pct*n)
n_train <- n - n_test
test_inds <- sort(sample(1:n, n_test, replace = FALSE))
train_inds <- (1:n)[!((1:n) %in% test_inds)]
X_test <- as.data.frame(X[test_inds,])
X_train <- as.data.frame(X[train_inds,])
y_test <- y[test_inds]
y_train <- y[train_inds]n_test = round(0.2 * n)
test_inds = rng.choice(n, n_test, replace=False)
train_inds = np.setdiff1d(np.arange(n), test_inds)
X_test = X[test_inds]
X_train = X[train_inds]
y_test = y[test_inds]
y_train = y[train_inds]Model Sampling
Sample a BART model with 10 GFR and 100 MCMC iterations
num_gfr <- 10
num_burnin <- 0
num_mcmc <- 100
general_params <- list(keep_gfr = T)
bart_model <- stochtree::bart(
X_train = X_train, y_train = y_train, X_test = X_test,
num_gfr = num_gfr, num_burnin = num_burnin, num_mcmc = num_mcmc,
general_params = general_params
)num_gfr = 10
num_burnin = 0
num_mcmc = 100
bart_model = BARTModel()
bart_model.sample(
X_train=X_train, y_train=y_train, X_test=X_test,
num_gfr=num_gfr, num_burnin=num_burnin, num_mcmc=num_mcmc,
general_params={"num_threads": 1, "keep_gfr": True},
)Model Inspection
Assess the global error variance traceplot and test set prediction quality
sigma2_samples <- extractParameter(bart_model, "sigma2_global")
plot(sigma2_samples, ylab="sigma^2")
abline(h=noise_sd^2,col="red",lty=2,lwd=2.5)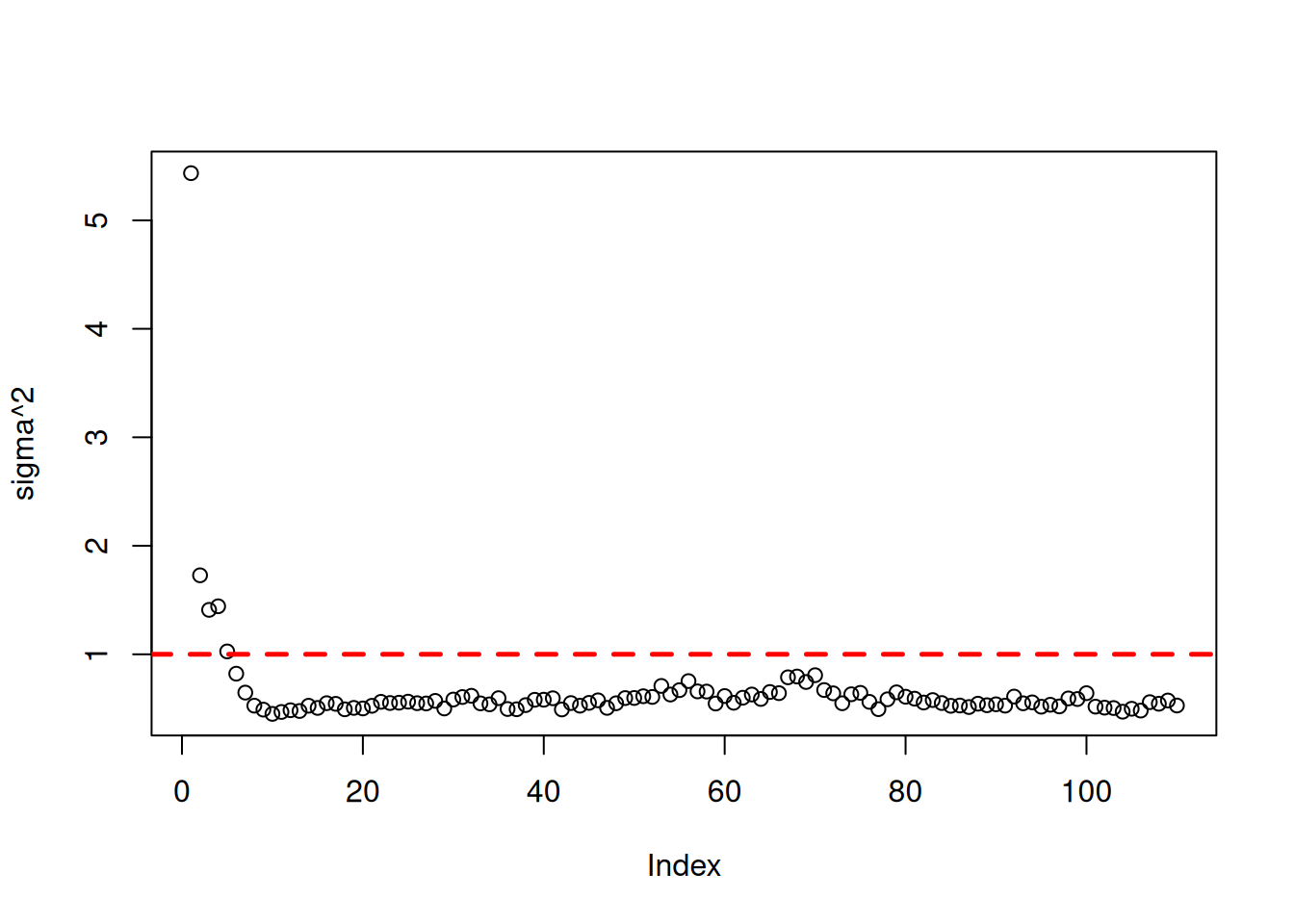
y_hat_test <- predict(bart_model, X=X_test, type="mean", terms="y_hat")
plot(y_hat_test, y_test, pch=16, cex=0.75, xlab = "pred", ylab = "actual")
abline(0,1,col="red",lty=2,lwd=2.5)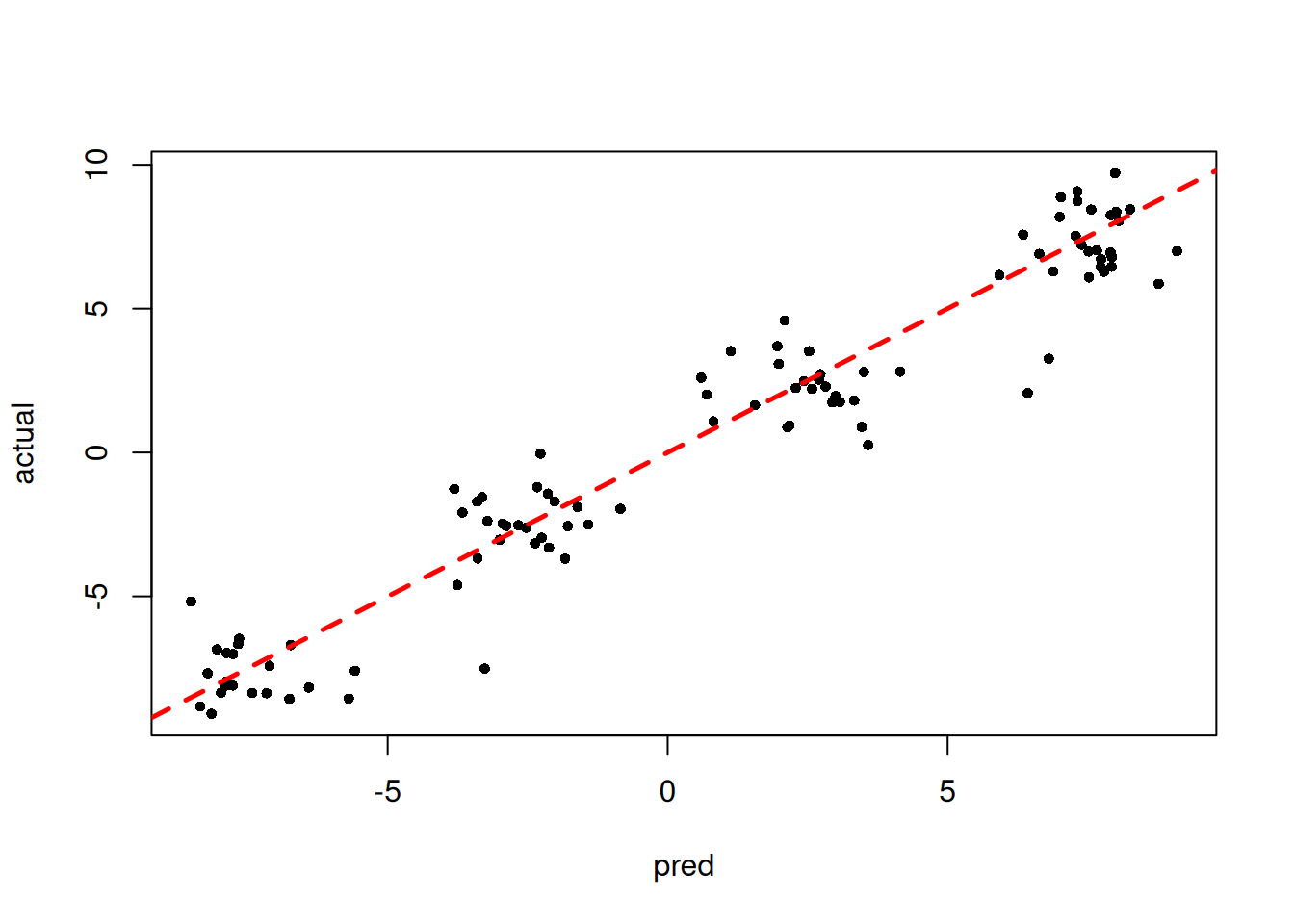
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
sigma2_samples = bart_model.extract_parameter("sigma2_global")
ax1.plot(sigma2_samples)
ax1.axhline(noise_sd**2, color="red", linestyle="dashed", linewidth=2)
ax1.set_ylabel(r"$\sigma^2$")
y_hat_test = bart_model.predict(X=X_test, terms="y_hat", type="mean")
ax2.scatter(y_hat_test, y_test, s=15, alpha=0.6)
lo = min(y_hat_test.min(), y_test.min())
hi = max(y_hat_test.max(), y_test.max())
ax2.plot([lo, hi], [lo, hi], color="red", linestyle="dashed", linewidth=2)
ax2.set_xlabel("pred")
ax2.set_ylabel("actual")
plt.tight_layout()
plt.show()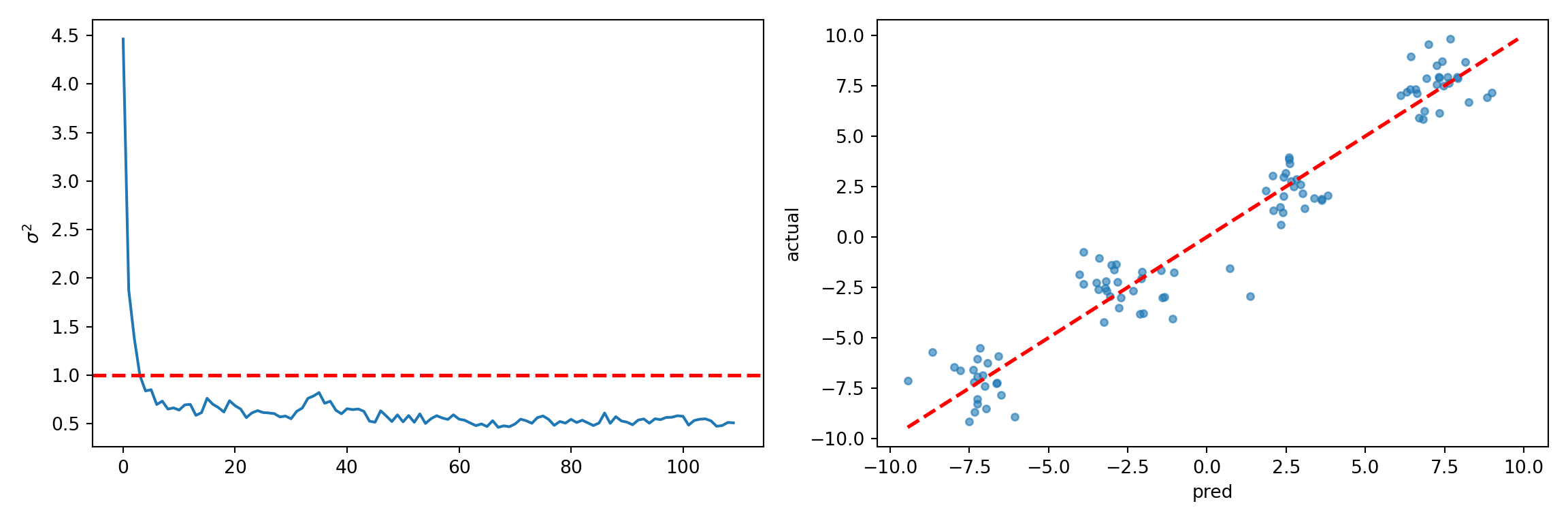
Variable Split Counts
The get_forest_split_counts method of a BART model’s internal forest objects allows us to compute the number of times each variable was used in a split rule across all trees in a given forest.
Below we query this vector for the final GFR sample (1-indexed as 10 in R, 0-indexed as 9 in Python), where the second argument is the dimensionality of the covariates.
bart_model$mean_forests$get_forest_split_counts(10, p_x) [1] 26 23 26 33 25 31 21 26 24 36bart_model.forest_container_mean.get_forest_split_counts(9, p_x)array([28, 32, 20, 36, 20, 29, 27, 22, 30, 32], dtype=int32)We can also compute split counts for each feature aggregated over all forests
bart_model$mean_forests$get_aggregate_split_counts(p_x) [1] 2472 3543 2643 2829 2978 2438 2130 2682 1892 3508bart_model.forest_container_mean.get_overall_split_counts(p_x)array([3085, 3253, 2030, 2975, 2141, 2676, 2514, 2471, 2586, 3393],
dtype=int32)The split counts appear relatively uniform across features, so let’s dig deeper and look at individual trees.
The get_granular_split_counts method returns a 3-dimensional array of shape (num_forests, num_trees, num_features), where each entry represents the number of times a feature was used in a split for a specific tree in a specific forest.
That is, we can count the number of times feature \(k\) was split on in tree \(j\) of forest \(i\) by looking at the (i,j,k) entry of this array.
Below we compute the split count for all features in the first tree of the last GFR sample in our model (noting again the use of 1-indexing in R and 0-indexing in Python).
splits = bart_model$mean_forests$get_granular_split_counts(p_x)
splits[10,1,] [1] 0 0 0 0 0 0 0 0 0 1splits = bart_model.forest_container_mean.get_granular_split_counts(p_x)
splits[9, 0, :]array([0, 0, 0, 0, 0, 0, 0, 0, 0, 1], dtype=int32)This tree has a single split on the only “important” feature (10). Now, let’s look at the second tree.
splits[10,2,] [1] 0 0 0 0 0 0 0 0 0 2splits[9, 1, :]array([0, 0, 0, 0, 0, 0, 0, 0, 0, 1], dtype=int32)And the 20th and 30th trees
splits[10,20,] [1] 0 0 0 0 0 1 0 0 0 0splits[9, 19, :]array([0, 0, 0, 0, 0, 0, 0, 0, 0, 2], dtype=int32)splits[10,30,] [1] 0 0 0 0 0 0 0 1 0 0splits[9, 29, :]array([0, 0, 0, 0, 0, 0, 0, 1, 0, 0], dtype=int32)We see that “later” trees are splitting on other features, but we also note that these trees are fitting an outcome that is already residualized by many “relevant splits” made by trees 1 and 2.
Tree Structure
Now, let’s inspect the first tree for the last GFR sample in more depth, following this scikit-learn vignette.
forest_num <- 9
tree_num <- 0
nodes <- sort(bart_model$mean_forests$nodes(forest_num, tree_num))
for (nid in nodes) {
if (bart_model$mean_forests$is_leaf_node(forest_num, tree_num, nid)) {
node_depth <- bart_model$mean_forests$node_depth(forest_num, tree_num, nid)
space_text <- rep("\t", node_depth)
leaf_values <- bart_model$mean_forests$node_leaf_values(forest_num, tree_num, nid)
cat(space_text, "node=", nid, " is a leaf node with value=",
format(leaf_values, digits = 3), "\n", sep = "")
} else {
node_depth <- bart_model$mean_forests$node_depth(forest_num, tree_num, nid)
space_text <- rep("\t", node_depth)
left <- bart_model$mean_forests$left_child_node(forest_num, tree_num, nid)
feature <- bart_model$mean_forests$node_split_index(forest_num, tree_num, nid)
threshold <- bart_model$mean_forests$node_split_threshold(forest_num, tree_num, nid)
right <- bart_model$mean_forests$right_child_node(forest_num, tree_num, nid)
cat(space_text, "node=", nid, " is a split node, which tells us to go to node ",
left, " if X[:, ", feature, "] <= ", format(threshold, digits = 3),
" else to node ", right, "\n", sep = "")
}
}node=0 is a split node, which tells us to go to node 1 if X[:, 9] <= 0.497 else to node 2
node=1 is a leaf node with value=-0.467
node=2 is a leaf node with value=0.331forest_num = 9
tree_num = 0
fc = bart_model.forest_container_mean
nodes = np.sort(fc.nodes(forest_num, tree_num))
for nid in nodes:
depth = fc.node_depth(forest_num, tree_num, nid)
indent = "\t" * depth
if fc.is_leaf_node(forest_num, tree_num, nid):
value = np.round(fc.node_leaf_values(forest_num, tree_num, nid), 3)
print(f"{indent}node={nid} is a leaf node with value={value}")
else:
left = fc.left_child_node(forest_num, tree_num, nid)
feature = fc.node_split_index(forest_num, tree_num, nid)
threshold = round(fc.node_split_threshold(forest_num, tree_num, nid), 3)
right = fc.right_child_node(forest_num, tree_num, nid)
print(f"{indent}node={nid} is a split node, which tells us to go to node "
f"{left} if X[:, {feature}] <= {threshold} else to node {right}")node=0 is a split node, which tells us to go to node 1 if X[:, 9] <= 0.488 else to node 2
node=1 is a leaf node with value=[-0.317]
node=2 is a leaf node with value=[0.325]